Jiella endophytica
Taxonomy: cellular organisms; Bacteria; Proteobacteria; Alphaproteobacteria; Hyphomicrobiales; Aurantimonadaceae; Jiella
Average proteome isoelectric point is 6.27
Get precalculated fractions of proteins
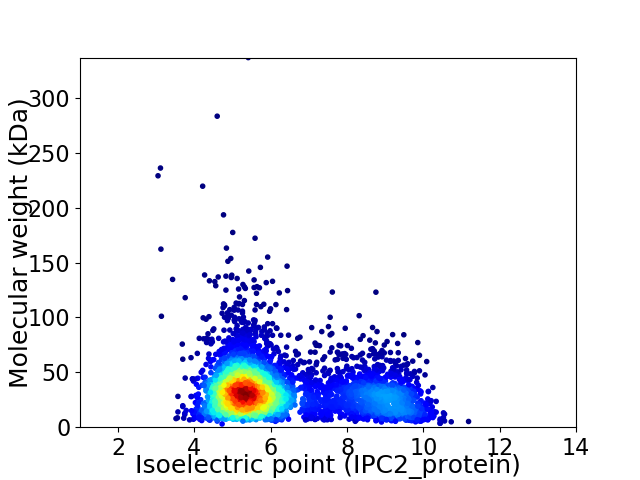
Virtual 2D-PAGE plot for 4547 proteins (isoelectric point calculated using IPC2_protein)
Get csv file with sequences according to given criteria:
* You can choose from 21 different methods for calculating isoelectric point
Summary statistics related to proteome-wise predictions
Protein with the lowest isoelectric point:
>tr|A0A4Y8R9K4|A0A4Y8R9K4_9RHIZ ABC transporter substrate-binding protein OS=Jiella endophytica OX=2558362 GN=E3C22_23485 PE=3 SV=1
MM1 pKa = 7.09IRR3 pKa = 11.84SRR5 pKa = 11.84FASLLFVGTSLAGMPALAQEE25 pKa = 4.98IPTSPPADD33 pKa = 3.75FSNGRR38 pKa = 11.84TPDD41 pKa = 3.89DD42 pKa = 4.62PSDD45 pKa = 3.65LPYY48 pKa = 11.5LMDD51 pKa = 4.45SYY53 pKa = 12.05VEE55 pKa = 4.56LLLDD59 pKa = 4.75DD60 pKa = 5.43PDD62 pKa = 5.3VMTQNYY68 pKa = 9.73DD69 pKa = 3.65YY70 pKa = 11.57VVTATQNRR78 pKa = 11.84TDD80 pKa = 3.91EE81 pKa = 4.12QTLAAIHH88 pKa = 7.48DD89 pKa = 4.2DD90 pKa = 3.6RR91 pKa = 11.84TNQQYY96 pKa = 11.42SVMNGLGVLTSIVMTGTGASADD118 pKa = 3.69GTTPASLTPTTYY130 pKa = 11.5ADD132 pKa = 3.37ATIADD137 pKa = 4.07YY138 pKa = 10.91EE139 pKa = 4.21ADD141 pKa = 3.27INYY144 pKa = 10.09LNGASWGTTTFADD157 pKa = 4.35GTATPLASAVTFIQDD172 pKa = 3.1IVRR175 pKa = 11.84ADD177 pKa = 3.49ASTEE181 pKa = 3.7PSKK184 pKa = 10.67RR185 pKa = 11.84TFDD188 pKa = 3.97RR189 pKa = 11.84LMEE192 pKa = 4.68NNDD195 pKa = 3.31TYY197 pKa = 11.08RR198 pKa = 11.84QYY200 pKa = 11.29EE201 pKa = 4.34SQYY204 pKa = 10.29QDD206 pKa = 3.14YY207 pKa = 10.93NVISNPDD214 pKa = 3.37ALTTEE219 pKa = 4.23DD220 pKa = 3.76TADD223 pKa = 4.56FIIPAYY229 pKa = 10.08LSNYY233 pKa = 9.17DD234 pKa = 3.35VPEE237 pKa = 4.25EE238 pKa = 4.1YY239 pKa = 9.64GTVEE243 pKa = 3.24NWVRR247 pKa = 11.84GFTVTQEE254 pKa = 4.3MIDD257 pKa = 3.77EE258 pKa = 4.64NGGEE262 pKa = 4.43PITVEE267 pKa = 5.11GLGLWTDD274 pKa = 4.04GVFTPATFGVGDD286 pKa = 4.01YY287 pKa = 11.06VPGIGTSPRR296 pKa = 11.84PYY298 pKa = 9.59RR299 pKa = 11.84VSDD302 pKa = 3.55NVAVPEE308 pKa = 4.52LLEE311 pKa = 4.14QVANTTNPYY320 pKa = 10.72ADD322 pKa = 3.14GSMPSGHH329 pKa = 6.49TSSAWTEE336 pKa = 3.85ALGLAFLVPQEE347 pKa = 4.11MQSLVTRR354 pKa = 11.84ASEE357 pKa = 4.03LGEE360 pKa = 3.94DD361 pKa = 3.7RR362 pKa = 11.84ILAGEE367 pKa = 4.44HH368 pKa = 6.37SPLDD372 pKa = 3.88VIGGRR377 pKa = 11.84IIAEE381 pKa = 4.71AIAATNIYY389 pKa = 9.77DD390 pKa = 4.54ALYY393 pKa = 10.44DD394 pKa = 4.09DD395 pKa = 4.78EE396 pKa = 5.71GNRR399 pKa = 11.84LDD401 pKa = 3.66WTDD404 pKa = 3.42PANEE408 pKa = 3.91EE409 pKa = 4.08AYY411 pKa = 10.16AVYY414 pKa = 8.61TAYY417 pKa = 9.78TEE419 pKa = 4.16TQAYY423 pKa = 8.44IAEE426 pKa = 4.5ACGTATVDD434 pKa = 3.11EE435 pKa = 4.79CVALGGDD442 pKa = 4.14GTSVAYY448 pKa = 8.32GTAEE452 pKa = 3.82ANKK455 pKa = 10.21AAFLYY460 pKa = 10.45RR461 pKa = 11.84MTYY464 pKa = 10.04GFEE467 pKa = 4.25ATSEE471 pKa = 4.37PYY473 pKa = 10.69RR474 pKa = 11.84MAVSDD479 pKa = 4.33VPVQAQVLLLTRR491 pKa = 11.84FPYY494 pKa = 9.62LTDD497 pKa = 3.46EE498 pKa = 4.15QRR500 pKa = 11.84LEE502 pKa = 4.22VLASTALDD510 pKa = 3.29ADD512 pKa = 4.01YY513 pKa = 10.96PLISGNTYY521 pKa = 10.79DD522 pKa = 4.86GWGKK526 pKa = 10.82LNLYY530 pKa = 9.83DD531 pKa = 4.27AADD534 pKa = 4.35GYY536 pKa = 11.72GYY538 pKa = 10.58FEE540 pKa = 5.48NDD542 pKa = 2.94VTVTMDD548 pKa = 3.8ASLGGYY554 pKa = 8.63NAADD558 pKa = 3.17TWANDD563 pKa = 2.99IAGPGGLVKK572 pKa = 10.81DD573 pKa = 4.04GTGKK577 pKa = 8.71LTLSGQDD584 pKa = 3.18TYY586 pKa = 11.64TGDD589 pKa = 3.44TVVAGGTLAIDD600 pKa = 4.06GSIVSNVRR608 pKa = 11.84VQRR611 pKa = 11.84RR612 pKa = 11.84AVLRR616 pKa = 11.84GVGVIGGGVTVEE628 pKa = 4.82DD629 pKa = 4.84GGTLAPGNSPGTLTVANSVTLSDD652 pKa = 3.52GAMSEE657 pKa = 4.01FDD659 pKa = 3.15IDD661 pKa = 3.84GTGTEE666 pKa = 4.47IGAGNYY672 pKa = 9.5SRR674 pKa = 11.84LEE676 pKa = 3.91LVGEE680 pKa = 4.22DD681 pKa = 4.81SIFTADD687 pKa = 4.05GEE689 pKa = 4.8LIPLLRR695 pKa = 11.84GITGDD700 pKa = 3.22ATNSYY705 pKa = 9.02TPDD708 pKa = 3.08IGTQFQIVVAEE719 pKa = 4.2GGVAGSYY726 pKa = 11.09DD727 pKa = 4.51SLTQPDD733 pKa = 4.02GLADD737 pKa = 3.47GTRR740 pKa = 11.84FDD742 pKa = 4.75TIYY745 pKa = 10.91GSNDD749 pKa = 2.55LRR751 pKa = 11.84LVVTPTAYY759 pKa = 10.09EE760 pKa = 4.34DD761 pKa = 3.81YY762 pKa = 10.6AASGNAANVAEE773 pKa = 4.59ALDD776 pKa = 4.19AGRR779 pKa = 11.84QAAGIRR785 pKa = 11.84LDD787 pKa = 4.76DD788 pKa = 4.95DD789 pKa = 3.98AASLYY794 pKa = 10.49DD795 pKa = 3.81ALYY798 pKa = 10.7ALSAADD804 pKa = 4.63LPAALGDD811 pKa = 4.47LAPQAYY817 pKa = 9.94GDD819 pKa = 3.87ALLVNRR825 pKa = 11.84DD826 pKa = 2.96SWYY829 pKa = 10.74LGSDD833 pKa = 3.07AVNRR837 pKa = 11.84AIEE840 pKa = 3.89ARR842 pKa = 11.84RR843 pKa = 11.84GGVVDD848 pKa = 3.78PTMQSAAQHH857 pKa = 6.2GMTAWLTALGQKK869 pKa = 10.52ASIDD873 pKa = 3.64SDD875 pKa = 3.82PASGYY880 pKa = 10.65DD881 pKa = 3.23QTLGGLIVGLDD892 pKa = 3.48GEE894 pKa = 4.59MGDD897 pKa = 4.34GFLLGASLGYY907 pKa = 10.98VNLDD911 pKa = 3.03ADD913 pKa = 4.87LDD915 pKa = 3.93QGASFSGDD923 pKa = 3.11SLQGRR928 pKa = 11.84IYY930 pKa = 10.13GSYY933 pKa = 9.59RR934 pKa = 11.84RR935 pKa = 11.84GIAFVDD941 pKa = 4.26GQFGFAYY948 pKa = 9.66TDD950 pKa = 3.8GEE952 pKa = 4.17VDD954 pKa = 3.05RR955 pKa = 11.84GVAGFGGMHH964 pKa = 7.11GDD966 pKa = 3.72ADD968 pKa = 3.89STSFGGEE975 pKa = 3.13IRR977 pKa = 11.84TGLRR981 pKa = 11.84YY982 pKa = 9.97GDD984 pKa = 3.93EE985 pKa = 4.2ATSFEE990 pKa = 4.33PSIALRR996 pKa = 11.84GVSVQLDD1003 pKa = 3.88GATEE1007 pKa = 4.05SGPAAVSFDD1016 pKa = 3.72DD1017 pKa = 4.26QSFDD1021 pKa = 3.5SLQSVLALGVSHH1033 pKa = 7.57RR1034 pKa = 11.84IALADD1039 pKa = 3.79GMSLTPAARR1048 pKa = 11.84IGWSHH1053 pKa = 6.39EE1054 pKa = 3.9FSDD1057 pKa = 3.6TDD1059 pKa = 3.88GAVTAAFVGAPGASFEE1075 pKa = 4.61TVGASIGRR1083 pKa = 11.84DD1084 pKa = 3.08AALVGVSAEE1093 pKa = 4.16FANGGAGSVFASYY1106 pKa = 10.62EE1107 pKa = 3.83GAFAEE1112 pKa = 4.45NQSSHH1117 pKa = 5.85SVTAGLKK1124 pKa = 10.01FSWW1127 pKa = 3.68
MM1 pKa = 7.09IRR3 pKa = 11.84SRR5 pKa = 11.84FASLLFVGTSLAGMPALAQEE25 pKa = 4.98IPTSPPADD33 pKa = 3.75FSNGRR38 pKa = 11.84TPDD41 pKa = 3.89DD42 pKa = 4.62PSDD45 pKa = 3.65LPYY48 pKa = 11.5LMDD51 pKa = 4.45SYY53 pKa = 12.05VEE55 pKa = 4.56LLLDD59 pKa = 4.75DD60 pKa = 5.43PDD62 pKa = 5.3VMTQNYY68 pKa = 9.73DD69 pKa = 3.65YY70 pKa = 11.57VVTATQNRR78 pKa = 11.84TDD80 pKa = 3.91EE81 pKa = 4.12QTLAAIHH88 pKa = 7.48DD89 pKa = 4.2DD90 pKa = 3.6RR91 pKa = 11.84TNQQYY96 pKa = 11.42SVMNGLGVLTSIVMTGTGASADD118 pKa = 3.69GTTPASLTPTTYY130 pKa = 11.5ADD132 pKa = 3.37ATIADD137 pKa = 4.07YY138 pKa = 10.91EE139 pKa = 4.21ADD141 pKa = 3.27INYY144 pKa = 10.09LNGASWGTTTFADD157 pKa = 4.35GTATPLASAVTFIQDD172 pKa = 3.1IVRR175 pKa = 11.84ADD177 pKa = 3.49ASTEE181 pKa = 3.7PSKK184 pKa = 10.67RR185 pKa = 11.84TFDD188 pKa = 3.97RR189 pKa = 11.84LMEE192 pKa = 4.68NNDD195 pKa = 3.31TYY197 pKa = 11.08RR198 pKa = 11.84QYY200 pKa = 11.29EE201 pKa = 4.34SQYY204 pKa = 10.29QDD206 pKa = 3.14YY207 pKa = 10.93NVISNPDD214 pKa = 3.37ALTTEE219 pKa = 4.23DD220 pKa = 3.76TADD223 pKa = 4.56FIIPAYY229 pKa = 10.08LSNYY233 pKa = 9.17DD234 pKa = 3.35VPEE237 pKa = 4.25EE238 pKa = 4.1YY239 pKa = 9.64GTVEE243 pKa = 3.24NWVRR247 pKa = 11.84GFTVTQEE254 pKa = 4.3MIDD257 pKa = 3.77EE258 pKa = 4.64NGGEE262 pKa = 4.43PITVEE267 pKa = 5.11GLGLWTDD274 pKa = 4.04GVFTPATFGVGDD286 pKa = 4.01YY287 pKa = 11.06VPGIGTSPRR296 pKa = 11.84PYY298 pKa = 9.59RR299 pKa = 11.84VSDD302 pKa = 3.55NVAVPEE308 pKa = 4.52LLEE311 pKa = 4.14QVANTTNPYY320 pKa = 10.72ADD322 pKa = 3.14GSMPSGHH329 pKa = 6.49TSSAWTEE336 pKa = 3.85ALGLAFLVPQEE347 pKa = 4.11MQSLVTRR354 pKa = 11.84ASEE357 pKa = 4.03LGEE360 pKa = 3.94DD361 pKa = 3.7RR362 pKa = 11.84ILAGEE367 pKa = 4.44HH368 pKa = 6.37SPLDD372 pKa = 3.88VIGGRR377 pKa = 11.84IIAEE381 pKa = 4.71AIAATNIYY389 pKa = 9.77DD390 pKa = 4.54ALYY393 pKa = 10.44DD394 pKa = 4.09DD395 pKa = 4.78EE396 pKa = 5.71GNRR399 pKa = 11.84LDD401 pKa = 3.66WTDD404 pKa = 3.42PANEE408 pKa = 3.91EE409 pKa = 4.08AYY411 pKa = 10.16AVYY414 pKa = 8.61TAYY417 pKa = 9.78TEE419 pKa = 4.16TQAYY423 pKa = 8.44IAEE426 pKa = 4.5ACGTATVDD434 pKa = 3.11EE435 pKa = 4.79CVALGGDD442 pKa = 4.14GTSVAYY448 pKa = 8.32GTAEE452 pKa = 3.82ANKK455 pKa = 10.21AAFLYY460 pKa = 10.45RR461 pKa = 11.84MTYY464 pKa = 10.04GFEE467 pKa = 4.25ATSEE471 pKa = 4.37PYY473 pKa = 10.69RR474 pKa = 11.84MAVSDD479 pKa = 4.33VPVQAQVLLLTRR491 pKa = 11.84FPYY494 pKa = 9.62LTDD497 pKa = 3.46EE498 pKa = 4.15QRR500 pKa = 11.84LEE502 pKa = 4.22VLASTALDD510 pKa = 3.29ADD512 pKa = 4.01YY513 pKa = 10.96PLISGNTYY521 pKa = 10.79DD522 pKa = 4.86GWGKK526 pKa = 10.82LNLYY530 pKa = 9.83DD531 pKa = 4.27AADD534 pKa = 4.35GYY536 pKa = 11.72GYY538 pKa = 10.58FEE540 pKa = 5.48NDD542 pKa = 2.94VTVTMDD548 pKa = 3.8ASLGGYY554 pKa = 8.63NAADD558 pKa = 3.17TWANDD563 pKa = 2.99IAGPGGLVKK572 pKa = 10.81DD573 pKa = 4.04GTGKK577 pKa = 8.71LTLSGQDD584 pKa = 3.18TYY586 pKa = 11.64TGDD589 pKa = 3.44TVVAGGTLAIDD600 pKa = 4.06GSIVSNVRR608 pKa = 11.84VQRR611 pKa = 11.84RR612 pKa = 11.84AVLRR616 pKa = 11.84GVGVIGGGVTVEE628 pKa = 4.82DD629 pKa = 4.84GGTLAPGNSPGTLTVANSVTLSDD652 pKa = 3.52GAMSEE657 pKa = 4.01FDD659 pKa = 3.15IDD661 pKa = 3.84GTGTEE666 pKa = 4.47IGAGNYY672 pKa = 9.5SRR674 pKa = 11.84LEE676 pKa = 3.91LVGEE680 pKa = 4.22DD681 pKa = 4.81SIFTADD687 pKa = 4.05GEE689 pKa = 4.8LIPLLRR695 pKa = 11.84GITGDD700 pKa = 3.22ATNSYY705 pKa = 9.02TPDD708 pKa = 3.08IGTQFQIVVAEE719 pKa = 4.2GGVAGSYY726 pKa = 11.09DD727 pKa = 4.51SLTQPDD733 pKa = 4.02GLADD737 pKa = 3.47GTRR740 pKa = 11.84FDD742 pKa = 4.75TIYY745 pKa = 10.91GSNDD749 pKa = 2.55LRR751 pKa = 11.84LVVTPTAYY759 pKa = 10.09EE760 pKa = 4.34DD761 pKa = 3.81YY762 pKa = 10.6AASGNAANVAEE773 pKa = 4.59ALDD776 pKa = 4.19AGRR779 pKa = 11.84QAAGIRR785 pKa = 11.84LDD787 pKa = 4.76DD788 pKa = 4.95DD789 pKa = 3.98AASLYY794 pKa = 10.49DD795 pKa = 3.81ALYY798 pKa = 10.7ALSAADD804 pKa = 4.63LPAALGDD811 pKa = 4.47LAPQAYY817 pKa = 9.94GDD819 pKa = 3.87ALLVNRR825 pKa = 11.84DD826 pKa = 2.96SWYY829 pKa = 10.74LGSDD833 pKa = 3.07AVNRR837 pKa = 11.84AIEE840 pKa = 3.89ARR842 pKa = 11.84RR843 pKa = 11.84GGVVDD848 pKa = 3.78PTMQSAAQHH857 pKa = 6.2GMTAWLTALGQKK869 pKa = 10.52ASIDD873 pKa = 3.64SDD875 pKa = 3.82PASGYY880 pKa = 10.65DD881 pKa = 3.23QTLGGLIVGLDD892 pKa = 3.48GEE894 pKa = 4.59MGDD897 pKa = 4.34GFLLGASLGYY907 pKa = 10.98VNLDD911 pKa = 3.03ADD913 pKa = 4.87LDD915 pKa = 3.93QGASFSGDD923 pKa = 3.11SLQGRR928 pKa = 11.84IYY930 pKa = 10.13GSYY933 pKa = 9.59RR934 pKa = 11.84RR935 pKa = 11.84GIAFVDD941 pKa = 4.26GQFGFAYY948 pKa = 9.66TDD950 pKa = 3.8GEE952 pKa = 4.17VDD954 pKa = 3.05RR955 pKa = 11.84GVAGFGGMHH964 pKa = 7.11GDD966 pKa = 3.72ADD968 pKa = 3.89STSFGGEE975 pKa = 3.13IRR977 pKa = 11.84TGLRR981 pKa = 11.84YY982 pKa = 9.97GDD984 pKa = 3.93EE985 pKa = 4.2ATSFEE990 pKa = 4.33PSIALRR996 pKa = 11.84GVSVQLDD1003 pKa = 3.88GATEE1007 pKa = 4.05SGPAAVSFDD1016 pKa = 3.72DD1017 pKa = 4.26QSFDD1021 pKa = 3.5SLQSVLALGVSHH1033 pKa = 7.57RR1034 pKa = 11.84IALADD1039 pKa = 3.79GMSLTPAARR1048 pKa = 11.84IGWSHH1053 pKa = 6.39EE1054 pKa = 3.9FSDD1057 pKa = 3.6TDD1059 pKa = 3.88GAVTAAFVGAPGASFEE1075 pKa = 4.61TVGASIGRR1083 pKa = 11.84DD1084 pKa = 3.08AALVGVSAEE1093 pKa = 4.16FANGGAGSVFASYY1106 pKa = 10.62EE1107 pKa = 3.83GAFAEE1112 pKa = 4.45NQSSHH1117 pKa = 5.85SVTAGLKK1124 pKa = 10.01FSWW1127 pKa = 3.68
Molecular weight: 118.01 kDa
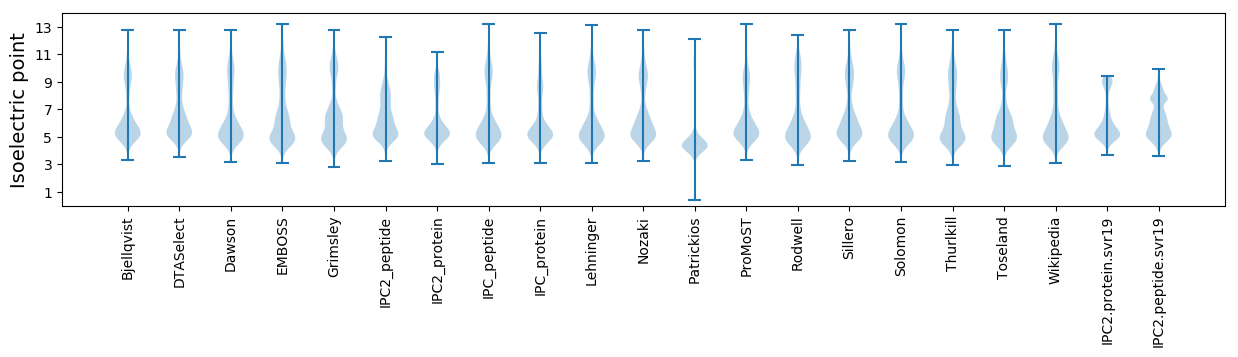
Isoelectric point according different methods:
Protein with the highest isoelectric point:
>tr|A0A4Y8RPI7|A0A4Y8RPI7_9RHIZ ABC transporter substrate-binding protein OS=Jiella endophytica OX=2558362 GN=E3C22_09120 PE=3 SV=1
MM1 pKa = 7.35KK2 pKa = 9.4RR3 pKa = 11.84TYY5 pKa = 10.2QPSRR9 pKa = 11.84LVRR12 pKa = 11.84KK13 pKa = 9.21RR14 pKa = 11.84RR15 pKa = 11.84HH16 pKa = 4.48GFRR19 pKa = 11.84ARR21 pKa = 11.84MATKK25 pKa = 10.38GGRR28 pKa = 11.84KK29 pKa = 9.14VIAARR34 pKa = 11.84RR35 pKa = 11.84SRR37 pKa = 11.84GRR39 pKa = 11.84KK40 pKa = 8.96RR41 pKa = 11.84LSAA44 pKa = 4.03
MM1 pKa = 7.35KK2 pKa = 9.4RR3 pKa = 11.84TYY5 pKa = 10.2QPSRR9 pKa = 11.84LVRR12 pKa = 11.84KK13 pKa = 9.21RR14 pKa = 11.84RR15 pKa = 11.84HH16 pKa = 4.48GFRR19 pKa = 11.84ARR21 pKa = 11.84MATKK25 pKa = 10.38GGRR28 pKa = 11.84KK29 pKa = 9.14VIAARR34 pKa = 11.84RR35 pKa = 11.84SRR37 pKa = 11.84GRR39 pKa = 11.84KK40 pKa = 8.96RR41 pKa = 11.84LSAA44 pKa = 4.03
Molecular weight: 5.21 kDa
Isoelectric point according different methods:
Peptides (in silico digests for buttom-up proteomics)
Below you can find in silico digests of the whole proteome with Trypsin, Chymotrypsin, Trypsin+LysC, LysN, ArgC proteases suitable for different mass spec machines.| Try ESI |
 |
|---|
| ChTry ESI |
 |
|---|
| ArgC ESI |
 |
|---|
| LysN ESI |
 |
|---|
| TryLysC ESI |
 |
|---|
| Try MALDI |
 |
|---|
| ChTry MALDI |
 |
|---|
| ArgC MALDI |
 |
|---|
| LysN MALDI |
 |
|---|
| TryLysC MALDI |
 |
|---|
| Try LTQ |
 |
|---|
| ChTry LTQ |
 |
|---|
| ArgC LTQ |
 |
|---|
| LysN LTQ |
 |
|---|
| TryLysC LTQ |
 |
|---|
| Try MSlow |
 |
|---|
| ChTry MSlow |
 |
|---|
| ArgC MSlow |
 |
|---|
| LysN MSlow |
 |
|---|
| TryLysC MSlow |
 |
|---|
| Try MShigh |
 |
|---|
| ChTry MShigh |
 |
|---|
| ArgC MShigh |
 |
|---|
| LysN MShigh |
 |
|---|
| TryLysC MShigh |
 |
|---|
General Statistics
Number of major isoforms |
Number of additional isoforms |
Number of all proteins |
Number of amino acids |
Min. Seq. Length |
Max. Seq. Length |
Avg. Seq. Length |
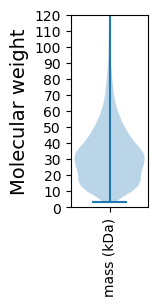
Avg. Mol. Weight |
|---|---|---|---|---|---|---|---|
0 |
1455190 |
25 |
3280 |
320.0 |
34.54 |
Amino acid frequency
Ala |
Cys |
Asp |
Glu |
Phe |
Gly |
His |
Ile |
Lys |
Leu |
|---|---|---|---|---|---|---|---|---|---|
13.024 ± 0.055 | 0.799 ± 0.01 |
5.88 ± 0.039 | 6.132 ± 0.037 |
3.828 ± 0.026 | 8.933 ± 0.04 |
1.861 ± 0.016 | 5.123 ± 0.03 |
3.18 ± 0.03 | 9.845 ± 0.048 |
Met |
Asn |
Gln |
Pro |
Arg |
Ser |
Thr |
Val |
Trp |
Tyr |
|---|---|---|---|---|---|---|---|---|---|
2.416 ± 0.018 | 2.329 ± 0.021 |
5.191 ± 0.033 | 2.734 ± 0.021 |
7.229 ± 0.04 | 5.55 ± 0.024 |
5.198 ± 0.029 | 7.429 ± 0.032 |
1.183 ± 0.013 | 2.134 ± 0.019 |
Most of the basic statistics you can see at this page can be downloaded from this CSV file
Proteome-pI is available under Creative Commons Attribution-NoDerivs license, for more details see here
| Reference: Kozlowski LP. Proteome-pI 2.0: Proteome Isoelectric Point Database Update. Nucleic Acids Res. 2021, doi: 10.1093/nar/gkab944 | Contact: Lukasz P. Kozlowski |
