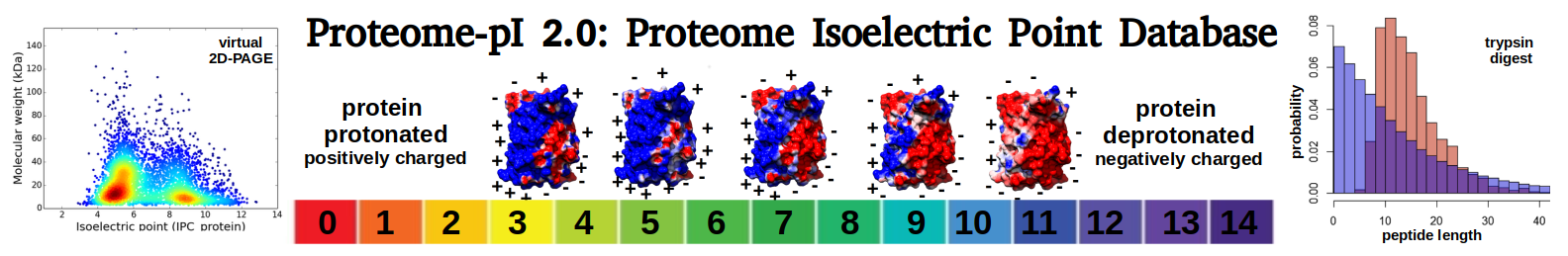
Bacillus phage Carmel_SA
Taxonomy: Viruses; Duplodnaviria; Heunggongvirae; Uroviricota; Caudoviricetes; Caudovirales; Siphoviridae; Wbetavirus; unclassified Wbetavirus
Average proteome isoelectric point is 6.73
Get precalculated fractions of proteins
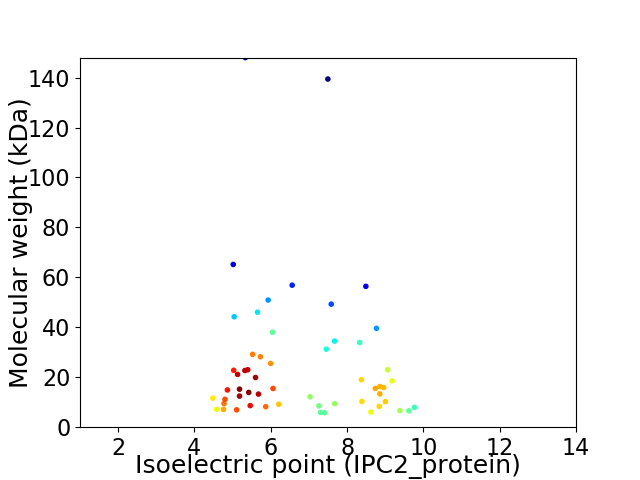
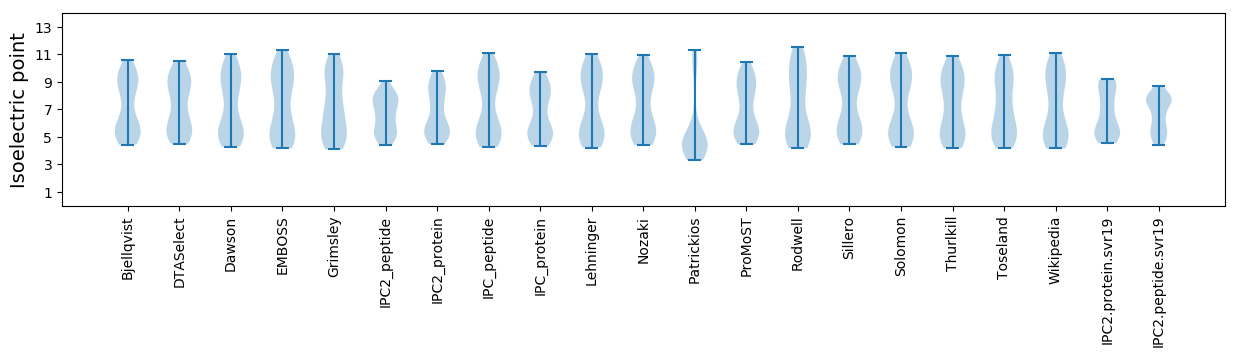
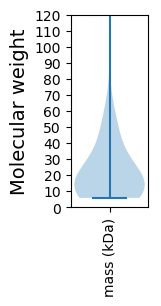
Virtual 2D-PAGE plot for 56 proteins (isoelectric point calculated using IPC2_protein)
Get csv file with sequences according to given criteria:
* You can choose from 21 different methods for calculating isoelectric point
Summary statistics related to proteome-wise predictions
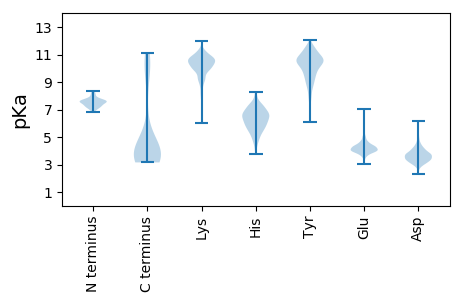
Protein with the lowest isoelectric point:
>tr|A0A288WGB0|A0A288WGB0_9CAUD Uncharacterized protein OS=Bacillus phage Carmel_SA OX=1983578 PE=4 SV=1
MM1 pKa = 7.71KK2 pKa = 9.33LTLRR6 pKa = 11.84INNEE10 pKa = 3.77EE11 pKa = 4.02KK12 pKa = 10.38TFNLPAFIPARR23 pKa = 11.84LIRR26 pKa = 11.84QAPEE30 pKa = 3.56LADD33 pKa = 3.2IPNNPEE39 pKa = 3.67AADD42 pKa = 3.96LDD44 pKa = 4.21KK45 pKa = 10.48MVQYY49 pKa = 10.47VVHH52 pKa = 6.61VYY54 pKa = 10.88GEE56 pKa = 4.19QFTVDD61 pKa = 3.81QYY63 pKa = 11.57WDD65 pKa = 3.51GVDD68 pKa = 3.15ARR70 pKa = 11.84KK71 pKa = 9.16FLLTTTNVINALVNEE86 pKa = 4.36TVEE89 pKa = 4.09AAGANPVADD98 pKa = 3.71GAEE101 pKa = 4.12NPNVV105 pKa = 3.33
MM1 pKa = 7.71KK2 pKa = 9.33LTLRR6 pKa = 11.84INNEE10 pKa = 3.77EE11 pKa = 4.02KK12 pKa = 10.38TFNLPAFIPARR23 pKa = 11.84LIRR26 pKa = 11.84QAPEE30 pKa = 3.56LADD33 pKa = 3.2IPNNPEE39 pKa = 3.67AADD42 pKa = 3.96LDD44 pKa = 4.21KK45 pKa = 10.48MVQYY49 pKa = 10.47VVHH52 pKa = 6.61VYY54 pKa = 10.88GEE56 pKa = 4.19QFTVDD61 pKa = 3.81QYY63 pKa = 11.57WDD65 pKa = 3.51GVDD68 pKa = 3.15ARR70 pKa = 11.84KK71 pKa = 9.16FLLTTTNVINALVNEE86 pKa = 4.36TVEE89 pKa = 4.09AAGANPVADD98 pKa = 3.71GAEE101 pKa = 4.12NPNVV105 pKa = 3.33
Molecular weight: 11.64 kDa
Isoelectric point according different methods:
Protein with the highest isoelectric point:
>tr|A0A288WG55|A0A288WG55_9CAUD Major tail protein OS=Bacillus phage Carmel_SA OX=1983578 PE=4 SV=1
MM1 pKa = 7.83CGNQNEE7 pKa = 4.74FIKK10 pKa = 10.72EE11 pKa = 3.83GSIKK15 pKa = 9.65MRR17 pKa = 11.84TWKK20 pKa = 10.2KK21 pKa = 9.78KK22 pKa = 8.18HH23 pKa = 5.82IKK25 pKa = 9.91RR26 pKa = 11.84AFLNRR31 pKa = 11.84QKK33 pKa = 10.83EE34 pKa = 4.15VDD36 pKa = 3.58KK37 pKa = 11.17EE38 pKa = 3.89RR39 pKa = 11.84TAAAWRR45 pKa = 11.84NIFVKK50 pKa = 10.61SGIIKK55 pKa = 10.38
MM1 pKa = 7.83CGNQNEE7 pKa = 4.74FIKK10 pKa = 10.72EE11 pKa = 3.83GSIKK15 pKa = 9.65MRR17 pKa = 11.84TWKK20 pKa = 10.2KK21 pKa = 9.78KK22 pKa = 8.18HH23 pKa = 5.82IKK25 pKa = 9.91RR26 pKa = 11.84AFLNRR31 pKa = 11.84QKK33 pKa = 10.83EE34 pKa = 4.15VDD36 pKa = 3.58KK37 pKa = 11.17EE38 pKa = 3.89RR39 pKa = 11.84TAAAWRR45 pKa = 11.84NIFVKK50 pKa = 10.61SGIIKK55 pKa = 10.38
Molecular weight: 6.56 kDa
Isoelectric point according different methods:
Peptides (in silico digests for buttom-up proteomics)
Below you can find in silico digests of the whole proteome with Trypsin, Chymotrypsin, Trypsin+LysC, LysN, ArgC proteases suitable for different mass spec machines.| Try ESI |
 |
|---|
| ChTry ESI |
 |
|---|
| ArgC ESI |
 |
|---|
| LysN ESI |
 |
|---|
| TryLysC ESI |
 |
|---|
| Try MALDI |
 |
|---|
| ChTry MALDI |
 |
|---|
| ArgC MALDI |
 |
|---|
| LysN MALDI |
 |
|---|
| TryLysC MALDI |
 |
|---|
| Try LTQ |
 |
|---|
| ChTry LTQ |
 |
|---|
| ArgC LTQ |
 |
|---|
| LysN LTQ |
 |
|---|
| TryLysC LTQ |
 |
|---|
| Try MSlow |
 |
|---|
| ChTry MSlow |
 |
|---|
| ArgC MSlow |
 |
|---|
| LysN MSlow |
 |
|---|
| TryLysC MSlow |
 |
|---|
| Try MShigh |
 |
|---|
| ChTry MShigh |
 |
|---|
| ArgC MShigh |
 |
|---|
| LysN MShigh |
 |
|---|
| TryLysC MShigh |
 |
|---|
General Statistics
Number of major isoforms |
Number of additional isoforms |
Number of all proteins |
Number of amino acids |
Min. Seq. Length |
Max. Seq. Length |
Avg. Seq. Length |
Avg. Mol. Weight |
|---|---|---|---|---|---|---|---|
0 |
12313 |
50 |
1321 |
219.9 |
25.19 |
Amino acid frequency
Ala |
Cys |
Asp |
Glu |
Phe |
Gly |
His |
Ile |
Lys |
Leu |
|---|---|---|---|---|---|---|---|---|---|
6.083 ± 0.584 | 0.926 ± 0.173 |
5.75 ± 0.202 | 8.56 ± 0.265 |
4.207 ± 0.228 | 5.685 ± 0.344 |
1.697 ± 0.153 | 6.741 ± 0.252 |
9.705 ± 0.327 | 8.601 ± 0.339 |
Met |
Asn |
Gln |
Pro |
Arg |
Ser |
Thr |
Val |
Trp |
Tyr |
|---|---|---|---|---|---|---|---|---|---|
2.631 ± 0.193 | 5.401 ± 0.212 |
2.631 ± 0.234 | 4.15 ± 0.276 |
4.621 ± 0.274 | 5.709 ± 0.213 |
5.588 ± 0.314 | 6.383 ± 0.229 |
1.121 ± 0.144 | 3.809 ± 0.27 |
Most of the basic statistics you can see at this page can be downloaded from this CSV file
Proteome-pI is available under Creative Commons Attribution-NoDerivs license, for more details see here
| Reference: Kozlowski LP. Proteome-pI 2.0: Proteome Isoelectric Point Database Update. Nucleic Acids Res. 2021, doi: 10.1093/nar/gkab944 | Contact: Lukasz P. Kozlowski |
