Sulfuricurvum sp. MLSB
Taxonomy: cellular organisms; Bacteria; Proteobacteria; delta/epsilon subdivisions; Epsilonproteobacteria; Campylobacterales; Thiovulaceae; Sulfuricurvum; unclassified Sulfuricurvum
Average proteome isoelectric point is 6.17
Get precalculated fractions of proteins
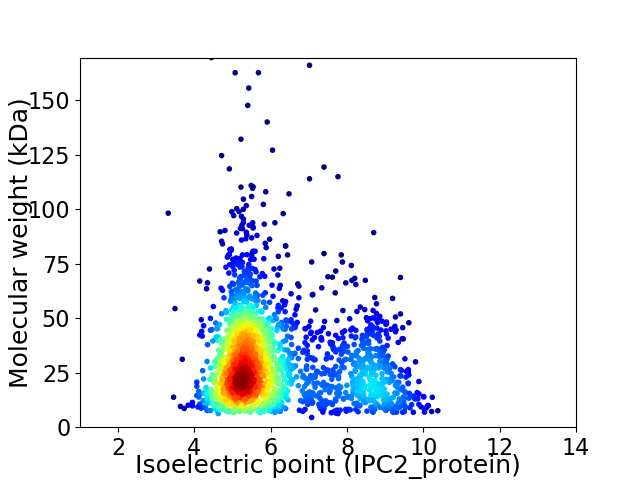
Virtual 2D-PAGE plot for 2005 proteins (isoelectric point calculated using IPC2_protein)
Get csv file with sequences according to given criteria:
* You can choose from 21 different methods for calculating isoelectric point
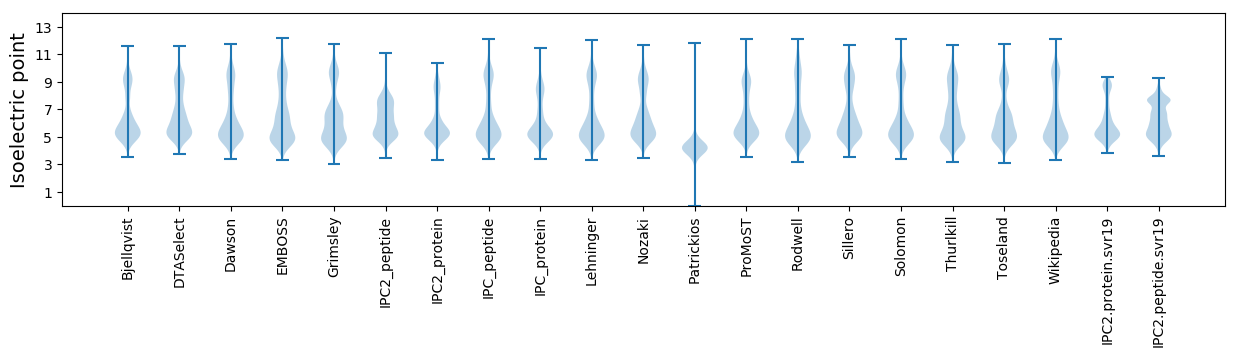
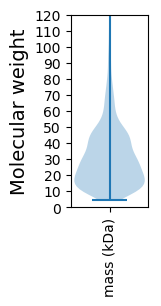
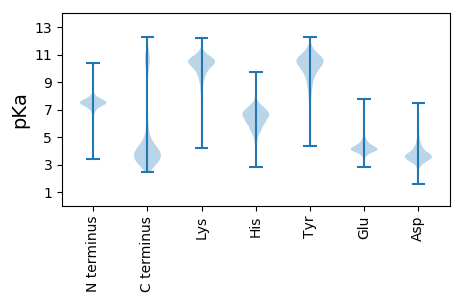
Summary statistics related to proteome-wise predictions
Protein with the lowest isoelectric point:
>tr|A0A091ANL1|A0A091ANL1_9PROT Protease HtpX homolog OS=Sulfuricurvum sp. MLSB OX=1537917 GN=htpX PE=3 SV=1
MM1 pKa = 7.23MNATKK6 pKa = 10.42LRR8 pKa = 11.84VMQAIFSLSDD18 pKa = 3.04EE19 pKa = 4.49TEE21 pKa = 4.13YY22 pKa = 11.36NIYY25 pKa = 10.25EE26 pKa = 4.22ADD28 pKa = 5.09DD29 pKa = 3.36IAEE32 pKa = 4.33YY33 pKa = 11.26ARR35 pKa = 11.84MDD37 pKa = 3.88ADD39 pKa = 3.45QVRR42 pKa = 11.84TIVSEE47 pKa = 4.39LYY49 pKa = 10.53DD50 pKa = 3.43EE51 pKa = 5.92GYY53 pKa = 10.68LGEE56 pKa = 4.72CMSIGDD62 pKa = 4.17DD63 pKa = 4.04GYY65 pKa = 10.7EE66 pKa = 3.73TFYY69 pKa = 11.32LNKK72 pKa = 9.93KK73 pKa = 8.88GRR75 pKa = 11.84ALIGMEE81 pKa = 3.93
MM1 pKa = 7.23MNATKK6 pKa = 10.42LRR8 pKa = 11.84VMQAIFSLSDD18 pKa = 3.04EE19 pKa = 4.49TEE21 pKa = 4.13YY22 pKa = 11.36NIYY25 pKa = 10.25EE26 pKa = 4.22ADD28 pKa = 5.09DD29 pKa = 3.36IAEE32 pKa = 4.33YY33 pKa = 11.26ARR35 pKa = 11.84MDD37 pKa = 3.88ADD39 pKa = 3.45QVRR42 pKa = 11.84TIVSEE47 pKa = 4.39LYY49 pKa = 10.53DD50 pKa = 3.43EE51 pKa = 5.92GYY53 pKa = 10.68LGEE56 pKa = 4.72CMSIGDD62 pKa = 4.17DD63 pKa = 4.04GYY65 pKa = 10.7EE66 pKa = 3.73TFYY69 pKa = 11.32LNKK72 pKa = 9.93KK73 pKa = 8.88GRR75 pKa = 11.84ALIGMEE81 pKa = 3.93
Molecular weight: 9.28 kDa
Isoelectric point according different methods:
Protein with the highest isoelectric point:
>tr|A0A091AM83|A0A091AM83_9PROT DNA polymerase III subunit gamma/tau OS=Sulfuricurvum sp. MLSB OX=1537917 GN=dnaX PE=3 SV=1
MM1 pKa = 7.21TKK3 pKa = 10.08SEE5 pKa = 4.11FRR7 pKa = 11.84SEE9 pKa = 3.94CTHH12 pKa = 6.49RR13 pKa = 11.84LKK15 pKa = 10.85NANRR19 pKa = 11.84STKK22 pKa = 10.19RR23 pKa = 11.84IRR25 pKa = 11.84DD26 pKa = 3.53AKK28 pKa = 8.81VRR30 pKa = 11.84RR31 pKa = 11.84MLQIVLKK38 pKa = 9.69EE39 pKa = 4.0LGPKK43 pKa = 9.58RR44 pKa = 11.84VLAFWPMEE52 pKa = 3.89FEE54 pKa = 3.96ADD56 pKa = 3.14IRR58 pKa = 11.84KK59 pKa = 9.65SLTTMRR65 pKa = 11.84RR66 pKa = 11.84WDD68 pKa = 3.56KK69 pKa = 11.27VYY71 pKa = 10.83LPFMDD76 pKa = 4.02GVSFKK81 pKa = 10.05MVPLRR86 pKa = 11.84LPLEE90 pKa = 4.25RR91 pKa = 11.84KK92 pKa = 9.81AFGIYY97 pKa = 9.43EE98 pKa = 4.12PRR100 pKa = 11.84DD101 pKa = 3.43SFRR104 pKa = 11.84KK105 pKa = 9.64INLIDD110 pKa = 3.66VAIVPAIGVDD120 pKa = 3.39ASLGRR125 pKa = 11.84IGFGKK130 pKa = 10.53GMYY133 pKa = 10.21DD134 pKa = 3.9RR135 pKa = 11.84FFPTLKK141 pKa = 9.52TKK143 pKa = 10.42PFTIFIQSCEE153 pKa = 3.79CRR155 pKa = 11.84TNRR158 pKa = 11.84FLCDD162 pKa = 3.93DD163 pKa = 3.82YY164 pKa = 11.86DD165 pKa = 3.75VRR167 pKa = 11.84SDD169 pKa = 4.93LLITPKK175 pKa = 10.5GVYY178 pKa = 9.61SMRR181 pKa = 11.84KK182 pKa = 5.42THH184 pKa = 6.0VKK186 pKa = 8.75RR187 pKa = 11.84TTRR190 pKa = 11.84GGRR193 pKa = 11.84HH194 pKa = 5.11SHH196 pKa = 6.14LQRR199 pKa = 11.84CSRR202 pKa = 11.84LFNFEE207 pKa = 3.41KK208 pKa = 10.68DD209 pKa = 3.5FRR211 pKa = 11.84RR212 pKa = 11.84SFF214 pKa = 3.32
MM1 pKa = 7.21TKK3 pKa = 10.08SEE5 pKa = 4.11FRR7 pKa = 11.84SEE9 pKa = 3.94CTHH12 pKa = 6.49RR13 pKa = 11.84LKK15 pKa = 10.85NANRR19 pKa = 11.84STKK22 pKa = 10.19RR23 pKa = 11.84IRR25 pKa = 11.84DD26 pKa = 3.53AKK28 pKa = 8.81VRR30 pKa = 11.84RR31 pKa = 11.84MLQIVLKK38 pKa = 9.69EE39 pKa = 4.0LGPKK43 pKa = 9.58RR44 pKa = 11.84VLAFWPMEE52 pKa = 3.89FEE54 pKa = 3.96ADD56 pKa = 3.14IRR58 pKa = 11.84KK59 pKa = 9.65SLTTMRR65 pKa = 11.84RR66 pKa = 11.84WDD68 pKa = 3.56KK69 pKa = 11.27VYY71 pKa = 10.83LPFMDD76 pKa = 4.02GVSFKK81 pKa = 10.05MVPLRR86 pKa = 11.84LPLEE90 pKa = 4.25RR91 pKa = 11.84KK92 pKa = 9.81AFGIYY97 pKa = 9.43EE98 pKa = 4.12PRR100 pKa = 11.84DD101 pKa = 3.43SFRR104 pKa = 11.84KK105 pKa = 9.64INLIDD110 pKa = 3.66VAIVPAIGVDD120 pKa = 3.39ASLGRR125 pKa = 11.84IGFGKK130 pKa = 10.53GMYY133 pKa = 10.21DD134 pKa = 3.9RR135 pKa = 11.84FFPTLKK141 pKa = 9.52TKK143 pKa = 10.42PFTIFIQSCEE153 pKa = 3.79CRR155 pKa = 11.84TNRR158 pKa = 11.84FLCDD162 pKa = 3.93DD163 pKa = 3.82YY164 pKa = 11.86DD165 pKa = 3.75VRR167 pKa = 11.84SDD169 pKa = 4.93LLITPKK175 pKa = 10.5GVYY178 pKa = 9.61SMRR181 pKa = 11.84KK182 pKa = 5.42THH184 pKa = 6.0VKK186 pKa = 8.75RR187 pKa = 11.84TTRR190 pKa = 11.84GGRR193 pKa = 11.84HH194 pKa = 5.11SHH196 pKa = 6.14LQRR199 pKa = 11.84CSRR202 pKa = 11.84LFNFEE207 pKa = 3.41KK208 pKa = 10.68DD209 pKa = 3.5FRR211 pKa = 11.84RR212 pKa = 11.84SFF214 pKa = 3.32
Molecular weight: 25.31 kDa
Isoelectric point according different methods:
Peptides (in silico digests for buttom-up proteomics)
Below you can find in silico digests of the whole proteome with Trypsin, Chymotrypsin, Trypsin+LysC, LysN, ArgC proteases suitable for different mass spec machines.| Try ESI |
 |
|---|
| ChTry ESI |
 |
|---|
| ArgC ESI |
 |
|---|
| LysN ESI |
 |
|---|
| TryLysC ESI |
 |
|---|
| Try MALDI |
 |
|---|
| ChTry MALDI |
 |
|---|
| ArgC MALDI |
 |
|---|
| LysN MALDI |
 |
|---|
| TryLysC MALDI |
 |
|---|
| Try LTQ |
 |
|---|
| ChTry LTQ |
 |
|---|
| ArgC LTQ |
 |
|---|
| LysN LTQ |
 |
|---|
| TryLysC LTQ |
 |
|---|
| Try MSlow |
 |
|---|
| ChTry MSlow |
 |
|---|
| ArgC MSlow |
 |
|---|
| LysN MSlow |
 |
|---|
| TryLysC MSlow |
 |
|---|
| Try MShigh |
 |
|---|
| ChTry MShigh |
 |
|---|
| ArgC MShigh |
 |
|---|
| LysN MShigh |
 |
|---|
| TryLysC MShigh |
 |
|---|
General Statistics
Number of major isoforms |
Number of additional isoforms |
Number of all proteins |
Number of amino acids |
Min. Seq. Length |
Max. Seq. Length |
Avg. Seq. Length |
Avg. Mol. Weight |
|---|---|---|---|---|---|---|---|
0 |
577772 |
39 |
1558 |
288.2 |
32.23 |
Amino acid frequency
Ala |
Cys |
Asp |
Glu |
Phe |
Gly |
His |
Ile |
Lys |
Leu |
|---|---|---|---|---|---|---|---|---|---|
8.426 ± 0.059 | 0.98 ± 0.024 |
5.529 ± 0.043 | 7.181 ± 0.059 |
4.513 ± 0.041 | 6.698 ± 0.051 |
2.307 ± 0.027 | 7.436 ± 0.054 |
6.211 ± 0.052 | 9.759 ± 0.06 |
Met |
Asn |
Gln |
Pro |
Arg |
Ser |
Thr |
Val |
Trp |
Tyr |
|---|---|---|---|---|---|---|---|---|---|
2.813 ± 0.029 | 4.146 ± 0.045 |
3.582 ± 0.032 | 3.076 ± 0.029 |
4.812 ± 0.039 | 6.266 ± 0.04 |
5.239 ± 0.045 | 6.549 ± 0.045 |
0.83 ± 0.019 | 3.647 ± 0.034 |
Most of the basic statistics you can see at this page can be downloaded from this CSV file
Proteome-pI is available under Creative Commons Attribution-NoDerivs license, for more details see here
| Reference: Kozlowski LP. Proteome-pI 2.0: Proteome Isoelectric Point Database Update. Nucleic Acids Res. 2021, doi: 10.1093/nar/gkab944 | Contact: Lukasz P. Kozlowski |
