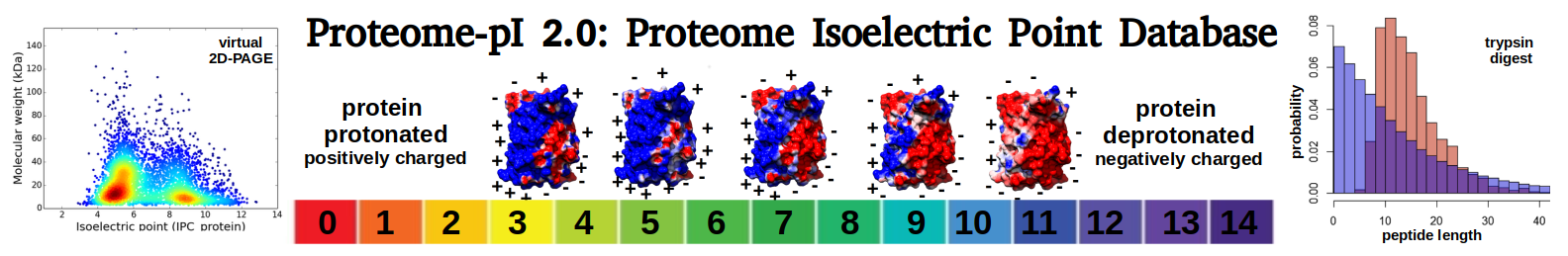
Streptococcus satellite phage Javan418
Taxonomy: Viruses; unclassified bacterial viruses
Average proteome isoelectric point is 6.63
Get precalculated fractions of proteins
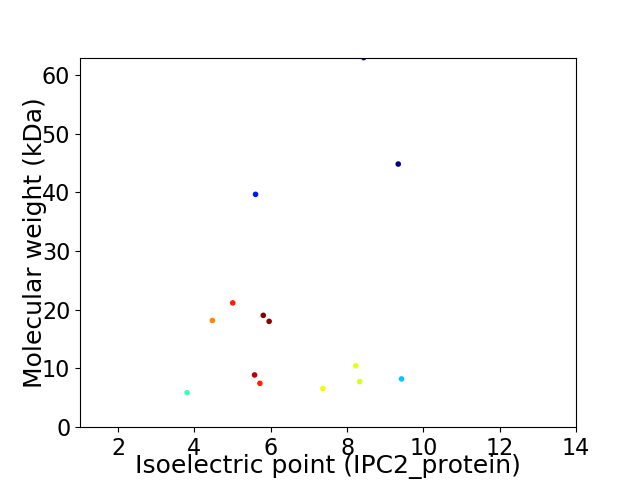
Virtual 2D-PAGE plot for 14 proteins (isoelectric point calculated using IPC2_protein)
Get csv file with sequences according to given criteria:
* You can choose from 21 different methods for calculating isoelectric point
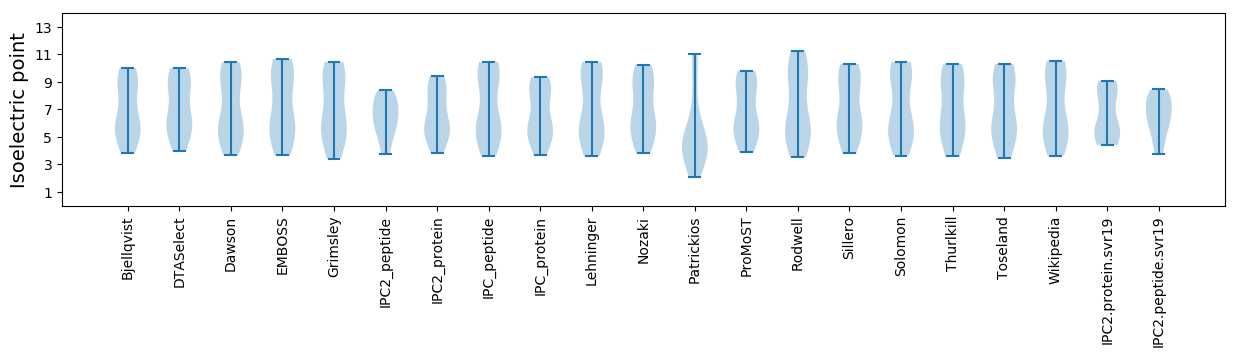
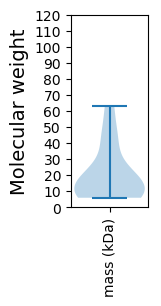
Summary statistics related to proteome-wise predictions
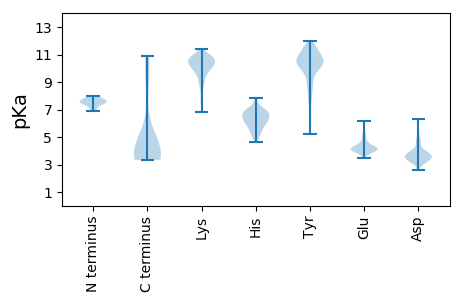
Protein with the lowest isoelectric point:
>tr|A0A4D5ZND1|A0A4D5ZND1_9VIRU Uncharacterized protein OS=Streptococcus satellite phage Javan418 OX=2558693 GN=JavanS418_0008 PE=4 SV=1
MM1 pKa = 7.74IDD3 pKa = 5.03DD4 pKa = 6.14IIPEE8 pKa = 4.39LNDD11 pKa = 3.55LQTLTTVALIDD22 pKa = 4.71DD23 pKa = 4.6NLNTSQAVLRR33 pKa = 11.84VIEE36 pKa = 4.06QKK38 pKa = 10.11IPRR41 pKa = 11.84LIEE44 pKa = 3.76ILEE47 pKa = 4.17EE48 pKa = 4.27LDD50 pKa = 3.45NGG52 pKa = 4.14
MM1 pKa = 7.74IDD3 pKa = 5.03DD4 pKa = 6.14IIPEE8 pKa = 4.39LNDD11 pKa = 3.55LQTLTTVALIDD22 pKa = 4.71DD23 pKa = 4.6NLNTSQAVLRR33 pKa = 11.84VIEE36 pKa = 4.06QKK38 pKa = 10.11IPRR41 pKa = 11.84LIEE44 pKa = 3.76ILEE47 pKa = 4.17EE48 pKa = 4.27LDD50 pKa = 3.45NGG52 pKa = 4.14
Molecular weight: 5.87 kDa
Isoelectric point according different methods:
Protein with the highest isoelectric point:
>tr|A0A4D5ZSH6|A0A4D5ZSH6_9VIRU DNA primase OS=Streptococcus satellite phage Javan418 OX=2558693 GN=JavanS418_0011 PE=4 SV=1
MM1 pKa = 7.51NIKK4 pKa = 10.04EE5 pKa = 4.29VIKK8 pKa = 10.83KK9 pKa = 10.43DD10 pKa = 3.21GTKK13 pKa = 9.85VYY15 pKa = 10.42RR16 pKa = 11.84SNVYY20 pKa = 10.53LGVDD24 pKa = 3.99SITGKK29 pKa = 10.34KK30 pKa = 10.21ARR32 pKa = 11.84TSVTARR38 pKa = 11.84TKK40 pKa = 10.93KK41 pKa = 9.82EE42 pKa = 3.91LKK44 pKa = 10.19QKK46 pKa = 10.6AKK48 pKa = 10.17QAITAFINNGYY59 pKa = 5.21TTKK62 pKa = 10.34TKK64 pKa = 10.73TNIRR68 pKa = 11.84TYY70 pKa = 11.15KK71 pKa = 9.7EE72 pKa = 3.7LAYY75 pKa = 10.49LWWDD79 pKa = 3.34SCKK82 pKa = 9.88NTVKK86 pKa = 10.52INTRR90 pKa = 11.84KK91 pKa = 9.89AVEE94 pKa = 3.95GQIRR98 pKa = 11.84VHH100 pKa = 6.67LLPAFGDD107 pKa = 3.81YY108 pKa = 10.62KK109 pKa = 10.88LEE111 pKa = 4.17KK112 pKa = 9.34LTTPIIQQQVNKK124 pKa = 8.81WANDD128 pKa = 3.35YY129 pKa = 11.53NKK131 pKa = 10.46GKK133 pKa = 10.19KK134 pKa = 9.96GSYY137 pKa = 8.57KK138 pKa = 10.37HH139 pKa = 6.14YY140 pKa = 10.4NQLHH144 pKa = 6.17ALNKK148 pKa = 10.23RR149 pKa = 11.84ILQYY153 pKa = 11.28AVIMQLIPFNPARR166 pKa = 11.84EE167 pKa = 4.22VIVPRR172 pKa = 11.84KK173 pKa = 9.67KK174 pKa = 8.87QTEE177 pKa = 4.14EE178 pKa = 3.63IKK180 pKa = 10.67IKK182 pKa = 10.62FLDD185 pKa = 3.76KK186 pKa = 11.2QEE188 pKa = 4.57LKK190 pKa = 10.68QFYY193 pKa = 10.2DD194 pKa = 3.74YY195 pKa = 11.63LDD197 pKa = 3.84TLDD200 pKa = 3.24KK201 pKa = 10.91SKK203 pKa = 11.21YY204 pKa = 8.38YY205 pKa = 11.15NLFDD209 pKa = 3.55VVLYY213 pKa = 8.25KK214 pKa = 10.5TLLATGCRR222 pKa = 11.84IGEE225 pKa = 4.15VLALEE230 pKa = 4.42WSDD233 pKa = 3.44IDD235 pKa = 5.15LDD237 pKa = 3.77NGYY240 pKa = 10.33ISINKK245 pKa = 7.82TLNRR249 pKa = 11.84DD250 pKa = 3.77DD251 pKa = 4.94EE252 pKa = 5.09VNCPKK257 pKa = 10.71SKK259 pKa = 10.67AGYY262 pKa = 9.69RR263 pKa = 11.84DD264 pKa = 3.13ISIDD268 pKa = 3.12KK269 pKa = 9.51ATILMLKK276 pKa = 9.61HH277 pKa = 5.3YY278 pKa = 10.2KK279 pKa = 9.78NRR281 pKa = 11.84QTIQAWQHH289 pKa = 5.19KK290 pKa = 8.15RR291 pKa = 11.84TEE293 pKa = 4.35TVVFSTFVNKK303 pKa = 10.04YY304 pKa = 9.36ASQPALRR311 pKa = 11.84RR312 pKa = 11.84RR313 pKa = 11.84LEE315 pKa = 3.89RR316 pKa = 11.84HH317 pKa = 5.84LKK319 pKa = 8.68NAGVSYY325 pKa = 10.82VAFHH329 pKa = 6.71GLRR332 pKa = 11.84HH333 pKa = 4.61THH335 pKa = 5.07ATIMLNAGIQPKK347 pKa = 10.11DD348 pKa = 3.26LQYY351 pKa = 11.63RR352 pKa = 11.84LGHH355 pKa = 6.02SNISMTLNTYY365 pKa = 8.1VHH367 pKa = 5.2VTKK370 pKa = 10.79EE371 pKa = 3.89GAKK374 pKa = 9.74NSASFFEE381 pKa = 4.46SAINSLGG388 pKa = 3.34
MM1 pKa = 7.51NIKK4 pKa = 10.04EE5 pKa = 4.29VIKK8 pKa = 10.83KK9 pKa = 10.43DD10 pKa = 3.21GTKK13 pKa = 9.85VYY15 pKa = 10.42RR16 pKa = 11.84SNVYY20 pKa = 10.53LGVDD24 pKa = 3.99SITGKK29 pKa = 10.34KK30 pKa = 10.21ARR32 pKa = 11.84TSVTARR38 pKa = 11.84TKK40 pKa = 10.93KK41 pKa = 9.82EE42 pKa = 3.91LKK44 pKa = 10.19QKK46 pKa = 10.6AKK48 pKa = 10.17QAITAFINNGYY59 pKa = 5.21TTKK62 pKa = 10.34TKK64 pKa = 10.73TNIRR68 pKa = 11.84TYY70 pKa = 11.15KK71 pKa = 9.7EE72 pKa = 3.7LAYY75 pKa = 10.49LWWDD79 pKa = 3.34SCKK82 pKa = 9.88NTVKK86 pKa = 10.52INTRR90 pKa = 11.84KK91 pKa = 9.89AVEE94 pKa = 3.95GQIRR98 pKa = 11.84VHH100 pKa = 6.67LLPAFGDD107 pKa = 3.81YY108 pKa = 10.62KK109 pKa = 10.88LEE111 pKa = 4.17KK112 pKa = 9.34LTTPIIQQQVNKK124 pKa = 8.81WANDD128 pKa = 3.35YY129 pKa = 11.53NKK131 pKa = 10.46GKK133 pKa = 10.19KK134 pKa = 9.96GSYY137 pKa = 8.57KK138 pKa = 10.37HH139 pKa = 6.14YY140 pKa = 10.4NQLHH144 pKa = 6.17ALNKK148 pKa = 10.23RR149 pKa = 11.84ILQYY153 pKa = 11.28AVIMQLIPFNPARR166 pKa = 11.84EE167 pKa = 4.22VIVPRR172 pKa = 11.84KK173 pKa = 9.67KK174 pKa = 8.87QTEE177 pKa = 4.14EE178 pKa = 3.63IKK180 pKa = 10.67IKK182 pKa = 10.62FLDD185 pKa = 3.76KK186 pKa = 11.2QEE188 pKa = 4.57LKK190 pKa = 10.68QFYY193 pKa = 10.2DD194 pKa = 3.74YY195 pKa = 11.63LDD197 pKa = 3.84TLDD200 pKa = 3.24KK201 pKa = 10.91SKK203 pKa = 11.21YY204 pKa = 8.38YY205 pKa = 11.15NLFDD209 pKa = 3.55VVLYY213 pKa = 8.25KK214 pKa = 10.5TLLATGCRR222 pKa = 11.84IGEE225 pKa = 4.15VLALEE230 pKa = 4.42WSDD233 pKa = 3.44IDD235 pKa = 5.15LDD237 pKa = 3.77NGYY240 pKa = 10.33ISINKK245 pKa = 7.82TLNRR249 pKa = 11.84DD250 pKa = 3.77DD251 pKa = 4.94EE252 pKa = 5.09VNCPKK257 pKa = 10.71SKK259 pKa = 10.67AGYY262 pKa = 9.69RR263 pKa = 11.84DD264 pKa = 3.13ISIDD268 pKa = 3.12KK269 pKa = 9.51ATILMLKK276 pKa = 9.61HH277 pKa = 5.3YY278 pKa = 10.2KK279 pKa = 9.78NRR281 pKa = 11.84QTIQAWQHH289 pKa = 5.19KK290 pKa = 8.15RR291 pKa = 11.84TEE293 pKa = 4.35TVVFSTFVNKK303 pKa = 10.04YY304 pKa = 9.36ASQPALRR311 pKa = 11.84RR312 pKa = 11.84RR313 pKa = 11.84LEE315 pKa = 3.89RR316 pKa = 11.84HH317 pKa = 5.84LKK319 pKa = 8.68NAGVSYY325 pKa = 10.82VAFHH329 pKa = 6.71GLRR332 pKa = 11.84HH333 pKa = 4.61THH335 pKa = 5.07ATIMLNAGIQPKK347 pKa = 10.11DD348 pKa = 3.26LQYY351 pKa = 11.63RR352 pKa = 11.84LGHH355 pKa = 6.02SNISMTLNTYY365 pKa = 8.1VHH367 pKa = 5.2VTKK370 pKa = 10.79EE371 pKa = 3.89GAKK374 pKa = 9.74NSASFFEE381 pKa = 4.46SAINSLGG388 pKa = 3.34
Molecular weight: 44.81 kDa
Isoelectric point according different methods:
Peptides (in silico digests for buttom-up proteomics)
Below you can find in silico digests of the whole proteome with Trypsin, Chymotrypsin, Trypsin+LysC, LysN, ArgC proteases suitable for different mass spec machines.| Try ESI |
 |
|---|
| ChTry ESI |
 |
|---|
| ArgC ESI |
 |
|---|
| LysN ESI |
 |
|---|
| TryLysC ESI |
 |
|---|
| Try MALDI |
 |
|---|
| ChTry MALDI |
 |
|---|
| ArgC MALDI |
 |
|---|
| LysN MALDI |
 |
|---|
| TryLysC MALDI |
 |
|---|
| Try LTQ |
 |
|---|
| ChTry LTQ |
 |
|---|
| ArgC LTQ |
 |
|---|
| LysN LTQ |
 |
|---|
| TryLysC LTQ |
 |
|---|
| Try MSlow |
 |
|---|
| ChTry MSlow |
 |
|---|
| ArgC MSlow |
 |
|---|
| LysN MSlow |
 |
|---|
| TryLysC MSlow |
 |
|---|
| Try MShigh |
 |
|---|
| ChTry MShigh |
 |
|---|
| ArgC MShigh |
 |
|---|
| LysN MShigh |
 |
|---|
| TryLysC MShigh |
 |
|---|
General Statistics
Number of major isoforms |
Number of additional isoforms |
Number of all proteins |
Number of amino acids |
Min. Seq. Length |
Max. Seq. Length |
Avg. Seq. Length |
Avg. Mol. Weight |
|---|---|---|---|---|---|---|---|
0 |
2402 |
52 |
545 |
171.6 |
19.93 |
Amino acid frequency
Ala |
Cys |
Asp |
Glu |
Phe |
Gly |
His |
Ile |
Lys |
Leu |
|---|---|---|---|---|---|---|---|---|---|
5.495 ± 0.424 | 0.5 ± 0.163 |
6.203 ± 0.503 | 7.286 ± 0.986 |
4.163 ± 0.416 | 4.288 ± 0.431 |
1.582 ± 0.311 | 7.702 ± 0.501 |
9.992 ± 0.555 | 9.908 ± 0.447 |
Met |
Asn |
Gln |
Pro |
Arg |
Ser |
Thr |
Val |
Trp |
Tyr |
|---|---|---|---|---|---|---|---|---|---|
2.248 ± 0.344 | 5.662 ± 0.413 |
3.122 ± 0.495 | 5.329 ± 0.488 |
4.038 ± 0.395 | 4.996 ± 0.352 |
6.495 ± 0.57 | 4.746 ± 0.268 |
1.041 ± 0.106 | 5.204 ± 0.362 |
Most of the basic statistics you can see at this page can be downloaded from this CSV file
Proteome-pI is available under Creative Commons Attribution-NoDerivs license, for more details see here
| Reference: Kozlowski LP. Proteome-pI 2.0: Proteome Isoelectric Point Database Update. Nucleic Acids Res. 2021, doi: 10.1093/nar/gkab944 | Contact: Lukasz P. Kozlowski |
