Bombus vosnesenskii
Taxonomy: cellular organisms; Eukaryota; Opisthokonta; Metazoa; Eumetazoa; Bilateria; Protostomia; Ecdysozoa; Panarthropoda; Arthropoda; Mandibulata; Pancrustacea; Hexapoda; Insecta; Dicondylia; Pterygota; Neoptera; Endopterygota; Hymenoptera; Apocrita; Aculeata; Apoidea; Apidae; Apinae; Bombini;
Average proteome isoelectric point is 6.74
Get precalculated fractions of proteins
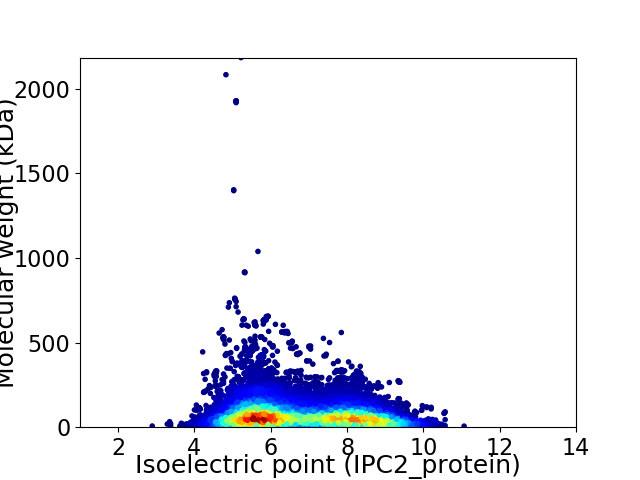
Virtual 2D-PAGE plot for 19375 proteins (isoelectric point calculated using IPC2_protein)
Get csv file with sequences according to given criteria:
* You can choose from 21 different methods for calculating isoelectric point
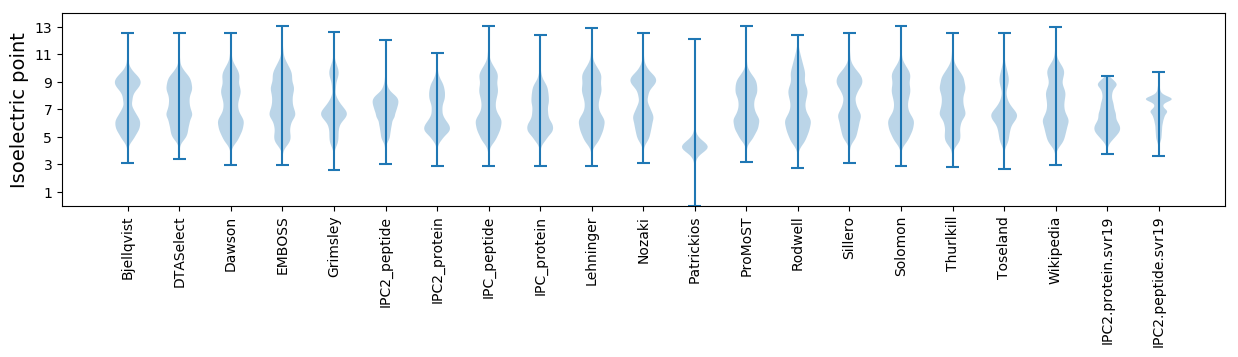
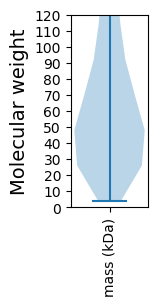
Summary statistics related to proteome-wise predictions
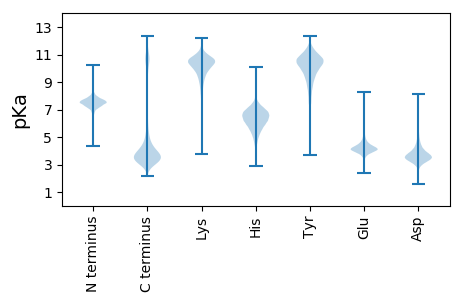
Protein with the lowest isoelectric point:
>tr|A0A6J3L2N8|A0A6J3L2N8_9HYME facilitated trehalose transporter Tret1-like isoform X1 OS=Bombus vosnesenskii OX=207650 GN=LOC117238603 PE=4 SV=1
MM1 pKa = 7.11VVLSNFLAPQEE12 pKa = 4.57GIRR15 pKa = 11.84HH16 pKa = 5.84EE17 pKa = 4.33EE18 pKa = 4.11LNTTVYY24 pKa = 10.75INDD27 pKa = 3.7RR28 pKa = 11.84EE29 pKa = 4.38VGKK32 pKa = 8.77GTLYY36 pKa = 9.3ITEE39 pKa = 4.71SLLSWVNYY47 pKa = 7.76DD48 pKa = 3.57TQQGFSLEE56 pKa = 4.16YY57 pKa = 10.02PHH59 pKa = 7.27ISLHH63 pKa = 6.32AISRR67 pKa = 11.84DD68 pKa = 3.61EE69 pKa = 3.96QAHH72 pKa = 6.04PRR74 pKa = 11.84QCLYY78 pKa = 10.88IMVDD82 pKa = 3.51AKK84 pKa = 11.27VDD86 pKa = 3.97LPDD89 pKa = 3.63VPLPPASDD97 pKa = 3.43SGSDD101 pKa = 4.18NEE103 pKa = 5.62FEE105 pKa = 5.5DD106 pKa = 4.37ADD108 pKa = 4.12TPITEE113 pKa = 4.27MRR115 pKa = 11.84FAPDD119 pKa = 3.05NTNNLEE125 pKa = 4.18TMFQAMNQCQALHH138 pKa = 7.44PDD140 pKa = 3.88PQDD143 pKa = 3.37SFSDD147 pKa = 3.74AEE149 pKa = 3.83EE150 pKa = 5.32DD151 pKa = 3.31IYY153 pKa = 11.51EE154 pKa = 4.54DD155 pKa = 4.26AEE157 pKa = 4.13EE158 pKa = 5.57DD159 pKa = 4.31DD160 pKa = 4.38FEE162 pKa = 5.13QYY164 pKa = 11.16DD165 pKa = 3.98VGAGDD170 pKa = 4.51APYY173 pKa = 10.34ILPTEE178 pKa = 4.35QIGTSHH184 pKa = 7.01NGTEE188 pKa = 4.35VDD190 pKa = 3.47DD191 pKa = 5.22AMDD194 pKa = 3.64IEE196 pKa = 4.94AGQFEE201 pKa = 4.76DD202 pKa = 5.49AEE204 pKa = 4.39EE205 pKa = 4.16DD206 pKa = 3.59LL207 pKa = 5.34
MM1 pKa = 7.11VVLSNFLAPQEE12 pKa = 4.57GIRR15 pKa = 11.84HH16 pKa = 5.84EE17 pKa = 4.33EE18 pKa = 4.11LNTTVYY24 pKa = 10.75INDD27 pKa = 3.7RR28 pKa = 11.84EE29 pKa = 4.38VGKK32 pKa = 8.77GTLYY36 pKa = 9.3ITEE39 pKa = 4.71SLLSWVNYY47 pKa = 7.76DD48 pKa = 3.57TQQGFSLEE56 pKa = 4.16YY57 pKa = 10.02PHH59 pKa = 7.27ISLHH63 pKa = 6.32AISRR67 pKa = 11.84DD68 pKa = 3.61EE69 pKa = 3.96QAHH72 pKa = 6.04PRR74 pKa = 11.84QCLYY78 pKa = 10.88IMVDD82 pKa = 3.51AKK84 pKa = 11.27VDD86 pKa = 3.97LPDD89 pKa = 3.63VPLPPASDD97 pKa = 3.43SGSDD101 pKa = 4.18NEE103 pKa = 5.62FEE105 pKa = 5.5DD106 pKa = 4.37ADD108 pKa = 4.12TPITEE113 pKa = 4.27MRR115 pKa = 11.84FAPDD119 pKa = 3.05NTNNLEE125 pKa = 4.18TMFQAMNQCQALHH138 pKa = 7.44PDD140 pKa = 3.88PQDD143 pKa = 3.37SFSDD147 pKa = 3.74AEE149 pKa = 3.83EE150 pKa = 5.32DD151 pKa = 3.31IYY153 pKa = 11.51EE154 pKa = 4.54DD155 pKa = 4.26AEE157 pKa = 4.13EE158 pKa = 5.57DD159 pKa = 4.31DD160 pKa = 4.38FEE162 pKa = 5.13QYY164 pKa = 11.16DD165 pKa = 3.98VGAGDD170 pKa = 4.51APYY173 pKa = 10.34ILPTEE178 pKa = 4.35QIGTSHH184 pKa = 7.01NGTEE188 pKa = 4.35VDD190 pKa = 3.47DD191 pKa = 5.22AMDD194 pKa = 3.64IEE196 pKa = 4.94AGQFEE201 pKa = 4.76DD202 pKa = 5.49AEE204 pKa = 4.39EE205 pKa = 4.16DD206 pKa = 3.59LL207 pKa = 5.34
Molecular weight: 23.23 kDa
Isoelectric point according different methods:
Protein with the highest isoelectric point:
>tr|A0A6J3KZR8|A0A6J3KZR8_9HYME NGFI-A-binding protein homolog isoform X1 OS=Bombus vosnesenskii OX=207650 GN=LOC117238230 PE=3 SV=1
MM1 pKa = 7.44SAHH4 pKa = 6.1KK5 pKa = 9.54TFIIKK10 pKa = 10.29RR11 pKa = 11.84KK12 pKa = 8.65LAKK15 pKa = 9.89KK16 pKa = 10.19LKK18 pKa = 8.64QNRR21 pKa = 11.84PIPQWVRR28 pKa = 11.84MRR30 pKa = 11.84TGNTIRR36 pKa = 11.84YY37 pKa = 5.79NAKK40 pKa = 8.47RR41 pKa = 11.84RR42 pKa = 11.84HH43 pKa = 4.1WRR45 pKa = 11.84RR46 pKa = 11.84TKK48 pKa = 10.89LKK50 pKa = 10.55LL51 pKa = 3.39
MM1 pKa = 7.44SAHH4 pKa = 6.1KK5 pKa = 9.54TFIIKK10 pKa = 10.29RR11 pKa = 11.84KK12 pKa = 8.65LAKK15 pKa = 9.89KK16 pKa = 10.19LKK18 pKa = 8.64QNRR21 pKa = 11.84PIPQWVRR28 pKa = 11.84MRR30 pKa = 11.84TGNTIRR36 pKa = 11.84YY37 pKa = 5.79NAKK40 pKa = 8.47RR41 pKa = 11.84RR42 pKa = 11.84HH43 pKa = 4.1WRR45 pKa = 11.84RR46 pKa = 11.84TKK48 pKa = 10.89LKK50 pKa = 10.55LL51 pKa = 3.39
Molecular weight: 6.36 kDa
Isoelectric point according different methods:
Peptides (in silico digests for buttom-up proteomics)
Below you can find in silico digests of the whole proteome with Trypsin, Chymotrypsin, Trypsin+LysC, LysN, ArgC proteases suitable for different mass spec machines.| Try ESI |
 |
|---|
| ChTry ESI |
 |
|---|
| ArgC ESI |
 |
|---|
| LysN ESI |
 |
|---|
| TryLysC ESI |
 |
|---|
| Try MALDI |
 |
|---|
| ChTry MALDI |
 |
|---|
| ArgC MALDI |
 |
|---|
| LysN MALDI |
 |
|---|
| TryLysC MALDI |
 |
|---|
| Try LTQ |
 |
|---|
| ChTry LTQ |
 |
|---|
| ArgC LTQ |
 |
|---|
| LysN LTQ |
 |
|---|
| TryLysC LTQ |
 |
|---|
| Try MSlow |
 |
|---|
| ChTry MSlow |
 |
|---|
| ArgC MSlow |
 |
|---|
| LysN MSlow |
 |
|---|
| TryLysC MSlow |
 |
|---|
| Try MShigh |
 |
|---|
| ChTry MShigh |
 |
|---|
| ArgC MShigh |
 |
|---|
| LysN MShigh |
 |
|---|
| TryLysC MShigh |
 |
|---|
General Statistics
Number of major isoforms |
Number of additional isoforms |
Number of all proteins |
Number of amino acids |
Min. Seq. Length |
Max. Seq. Length |
Avg. Seq. Length |
Avg. Mol. Weight |
|---|---|---|---|---|---|---|---|
14450676 |
34 |
19554 |
745.8 |
83.85 |
Amino acid frequency
Ala |
Cys |
Asp |
Glu |
Phe |
Gly |
His |
Ile |
Lys |
Leu |
|---|---|---|---|---|---|---|---|---|---|
6.093 ± 0.018 | 1.856 ± 0.02 |
5.373 ± 0.012 | 7.111 ± 0.033 |
3.288 ± 0.013 | 5.474 ± 0.02 |
2.493 ± 0.009 | 5.586 ± 0.015 |
6.368 ± 0.025 | 8.841 ± 0.023 |
Met |
Asn |
Gln |
Pro |
Arg |
Ser |
Thr |
Val |
Trp |
Tyr |
|---|---|---|---|---|---|---|---|---|---|
2.214 ± 0.007 | 5.112 ± 0.017 |
5.233 ± 0.024 | 4.688 ± 0.021 |
5.618 ± 0.017 | 8.513 ± 0.022 |
6.195 ± 0.019 | 5.956 ± 0.015 |
1.037 ± 0.006 | 2.95 ± 0.012 |
Most of the basic statistics you can see at this page can be downloaded from this CSV file
Proteome-pI is available under Creative Commons Attribution-NoDerivs license, for more details see here
| Reference: Kozlowski LP. Proteome-pI 2.0: Proteome Isoelectric Point Database Update. Nucleic Acids Res. 2021, doi: 10.1093/nar/gkab944 | Contact: Lukasz P. Kozlowski |
