Ursus americanus (American black bear) (Euarctos americanus)
Taxonomy: cellular organisms; Eukaryota; Opisthokonta; Metazoa; Eumetazoa; Bilateria; Deuterostomia; Chordata; Craniata; Vertebrata; Gnathostomata; Teleostomi; Euteleostomi; Sarcopterygii; Dipnotetrapodomorpha; Tetrapoda; Amniota; Mammalia; Theria; Eutheria; Boreoeutheria; Laurasiatheria;
Average proteome isoelectric point is 6.76
Get precalculated fractions of proteins
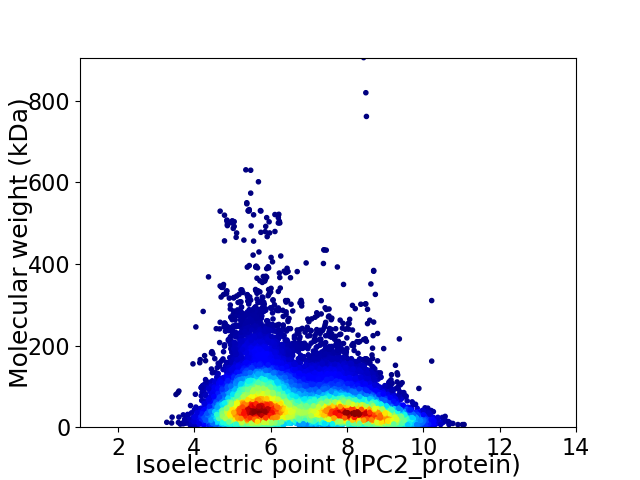
Virtual 2D-PAGE plot for 32512 proteins (isoelectric point calculated using IPC2_protein)
Get csv file with sequences according to given criteria:
* You can choose from 21 different methods for calculating isoelectric point
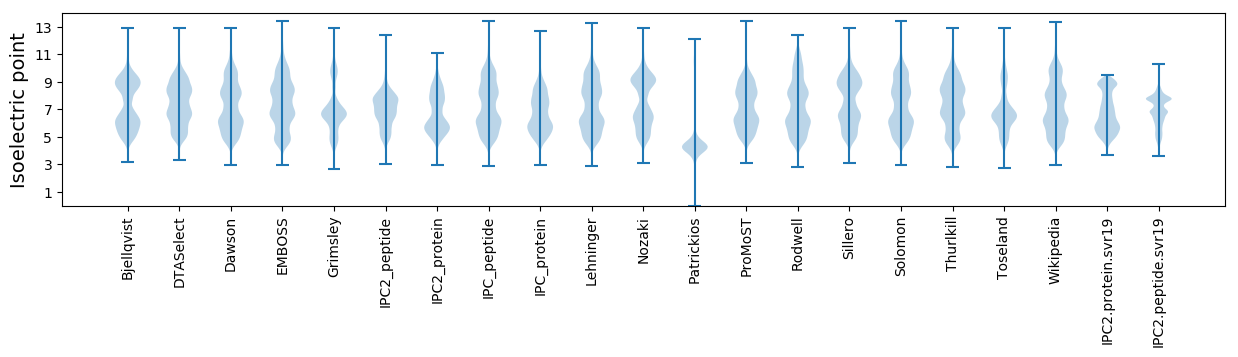
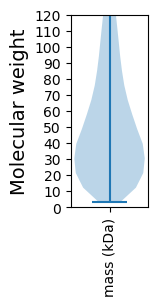
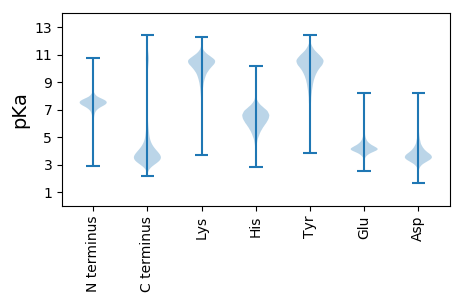
Summary statistics related to proteome-wise predictions
Protein with the lowest isoelectric point:
>tr|A0A452RLE4|A0A452RLE4_URSAM Isoform of A0A452RLJ5 Peptidyl-prolyl cis-trans isomerase OS=Ursus americanus OX=9643 GN=PPIC PE=3 SV=1
MM1 pKa = 7.29EE2 pKa = 4.36VVVLIPYY9 pKa = 10.0VLGLLLLPLLAVALCVRR26 pKa = 11.84CRR28 pKa = 11.84EE29 pKa = 4.18MPGSYY34 pKa = 8.87DD35 pKa = 3.14TGASDD40 pKa = 4.59SLTPSIMLKK49 pKa = 10.24RR50 pKa = 11.84PPTVVPWPSATSYY63 pKa = 11.6APVTSYY69 pKa = 11.21PPLSQPDD76 pKa = 4.23LLPIPRR82 pKa = 11.84SPQPHH87 pKa = 6.67RR88 pKa = 11.84MPSSRR93 pKa = 11.84QDD95 pKa = 2.92SDD97 pKa = 3.47GANSVASYY105 pKa = 10.71EE106 pKa = 4.2NEE108 pKa = 3.86EE109 pKa = 4.31PACEE113 pKa = 5.26DD114 pKa = 3.82DD115 pKa = 6.45DD116 pKa = 5.13EE117 pKa = 7.07DD118 pKa = 4.93EE119 pKa = 4.31EE120 pKa = 5.84DD121 pKa = 3.83YY122 pKa = 11.55HH123 pKa = 8.47NEE125 pKa = 4.01GYY127 pKa = 11.01LEE129 pKa = 4.22VLPDD133 pKa = 3.67STPATSTAVAPAPVPSNPGLRR154 pKa = 11.84DD155 pKa = 3.14SAFSMEE161 pKa = 4.23SGEE164 pKa = 4.92DD165 pKa = 3.63YY166 pKa = 11.71VNVAEE171 pKa = 4.65SEE173 pKa = 4.22EE174 pKa = 4.29SADD177 pKa = 3.6VSLDD181 pKa = 3.29GSRR184 pKa = 11.84EE185 pKa = 3.96YY186 pKa = 11.72VNVSQEE192 pKa = 4.44LPPMARR198 pKa = 11.84TEE200 pKa = 3.92AAILNSQKK208 pKa = 10.22VEE210 pKa = 4.2DD211 pKa = 4.43EE212 pKa = 4.21DD213 pKa = 5.59DD214 pKa = 3.99EE215 pKa = 4.77EE216 pKa = 5.11EE217 pKa = 4.32EE218 pKa = 4.72GAPDD222 pKa = 3.77YY223 pKa = 11.52EE224 pKa = 4.36NLQGLHH230 pKa = 6.14
MM1 pKa = 7.29EE2 pKa = 4.36VVVLIPYY9 pKa = 10.0VLGLLLLPLLAVALCVRR26 pKa = 11.84CRR28 pKa = 11.84EE29 pKa = 4.18MPGSYY34 pKa = 8.87DD35 pKa = 3.14TGASDD40 pKa = 4.59SLTPSIMLKK49 pKa = 10.24RR50 pKa = 11.84PPTVVPWPSATSYY63 pKa = 11.6APVTSYY69 pKa = 11.21PPLSQPDD76 pKa = 4.23LLPIPRR82 pKa = 11.84SPQPHH87 pKa = 6.67RR88 pKa = 11.84MPSSRR93 pKa = 11.84QDD95 pKa = 2.92SDD97 pKa = 3.47GANSVASYY105 pKa = 10.71EE106 pKa = 4.2NEE108 pKa = 3.86EE109 pKa = 4.31PACEE113 pKa = 5.26DD114 pKa = 3.82DD115 pKa = 6.45DD116 pKa = 5.13EE117 pKa = 7.07DD118 pKa = 4.93EE119 pKa = 4.31EE120 pKa = 5.84DD121 pKa = 3.83YY122 pKa = 11.55HH123 pKa = 8.47NEE125 pKa = 4.01GYY127 pKa = 11.01LEE129 pKa = 4.22VLPDD133 pKa = 3.67STPATSTAVAPAPVPSNPGLRR154 pKa = 11.84DD155 pKa = 3.14SAFSMEE161 pKa = 4.23SGEE164 pKa = 4.92DD165 pKa = 3.63YY166 pKa = 11.71VNVAEE171 pKa = 4.65SEE173 pKa = 4.22EE174 pKa = 4.29SADD177 pKa = 3.6VSLDD181 pKa = 3.29GSRR184 pKa = 11.84EE185 pKa = 3.96YY186 pKa = 11.72VNVSQEE192 pKa = 4.44LPPMARR198 pKa = 11.84TEE200 pKa = 3.92AAILNSQKK208 pKa = 10.22VEE210 pKa = 4.2DD211 pKa = 4.43EE212 pKa = 4.21DD213 pKa = 5.59DD214 pKa = 3.99EE215 pKa = 4.77EE216 pKa = 5.11EE217 pKa = 4.32EE218 pKa = 4.72GAPDD222 pKa = 3.77YY223 pKa = 11.52EE224 pKa = 4.36NLQGLHH230 pKa = 6.14
Molecular weight: 24.92 kDa
Isoelectric point according different methods:
Protein with the highest isoelectric point:
>tr|A0A452QCH8|A0A452QCH8_URSAM Coiled-coil domain containing 148 OS=Ursus americanus OX=9643 GN=CCDC148 PE=4 SV=1
PP1 pKa = 7.66ASPAGAPSLGASRR14 pKa = 11.84RR15 pKa = 11.84AAPRR19 pKa = 11.84WSAATPRR26 pKa = 11.84RR27 pKa = 11.84TTTTAAPSPPTPGAPRR43 pKa = 11.84RR44 pKa = 11.84PATRR48 pKa = 11.84SPPPRR53 pKa = 11.84LTTTTSSPPRR63 pKa = 11.84GARR66 pKa = 11.84RR67 pKa = 11.84AARR70 pKa = 11.84AARR73 pKa = 11.84AASRR77 pKa = 11.84ATRR80 pKa = 11.84ATARR84 pKa = 11.84RR85 pKa = 11.84RR86 pKa = 11.84RR87 pKa = 11.84TTANTAPRR95 pKa = 11.84RR96 pKa = 11.84PRR98 pKa = 11.84RR99 pKa = 11.84PWGTAATRR107 pKa = 11.84PSTRR111 pKa = 11.84SASTSPRR118 pKa = 11.84PGRR121 pKa = 11.84CPATRR126 pKa = 11.84CPSKK130 pKa = 10.47RR131 pKa = 11.84DD132 pKa = 3.36ARR134 pKa = 11.84RR135 pKa = 11.84SPPRR139 pKa = 11.84ATTRR143 pKa = 11.84RR144 pKa = 3.62
PP1 pKa = 7.66ASPAGAPSLGASRR14 pKa = 11.84RR15 pKa = 11.84AAPRR19 pKa = 11.84WSAATPRR26 pKa = 11.84RR27 pKa = 11.84TTTTAAPSPPTPGAPRR43 pKa = 11.84RR44 pKa = 11.84PATRR48 pKa = 11.84SPPPRR53 pKa = 11.84LTTTTSSPPRR63 pKa = 11.84GARR66 pKa = 11.84RR67 pKa = 11.84AARR70 pKa = 11.84AARR73 pKa = 11.84AASRR77 pKa = 11.84ATRR80 pKa = 11.84ATARR84 pKa = 11.84RR85 pKa = 11.84RR86 pKa = 11.84RR87 pKa = 11.84TTANTAPRR95 pKa = 11.84RR96 pKa = 11.84PRR98 pKa = 11.84RR99 pKa = 11.84PWGTAATRR107 pKa = 11.84PSTRR111 pKa = 11.84SASTSPRR118 pKa = 11.84PGRR121 pKa = 11.84CPATRR126 pKa = 11.84CPSKK130 pKa = 10.47RR131 pKa = 11.84DD132 pKa = 3.36ARR134 pKa = 11.84RR135 pKa = 11.84SPPRR139 pKa = 11.84ATTRR143 pKa = 11.84RR144 pKa = 3.62
Molecular weight: 15.28 kDa
Isoelectric point according different methods:
Peptides (in silico digests for buttom-up proteomics)
Below you can find in silico digests of the whole proteome with Trypsin, Chymotrypsin, Trypsin+LysC, LysN, ArgC proteases suitable for different mass spec machines.| Try ESI |
 |
|---|
| ChTry ESI |
 |
|---|
| ArgC ESI |
 |
|---|
| LysN ESI |
 |
|---|
| TryLysC ESI |
 |
|---|
| Try MALDI |
 |
|---|
| ChTry MALDI |
 |
|---|
| ArgC MALDI |
 |
|---|
| LysN MALDI |
 |
|---|
| TryLysC MALDI |
 |
|---|
| Try LTQ |
 |
|---|
| ChTry LTQ |
 |
|---|
| ArgC LTQ |
 |
|---|
| LysN LTQ |
 |
|---|
| TryLysC LTQ |
 |
|---|
| Try MSlow |
 |
|---|
| ChTry MSlow |
 |
|---|
| ArgC MSlow |
 |
|---|
| LysN MSlow |
 |
|---|
| TryLysC MSlow |
 |
|---|
| Try MShigh |
 |
|---|
| ChTry MShigh |
 |
|---|
| ArgC MShigh |
 |
|---|
| LysN MShigh |
 |
|---|
| TryLysC MShigh |
 |
|---|
General Statistics
Number of major isoforms |
Number of additional isoforms |
Number of all proteins |
Number of amino acids |
Min. Seq. Length |
Max. Seq. Length |
Avg. Seq. Length |
Avg. Mol. Weight |
|---|---|---|---|---|---|---|---|
18648622 |
27 |
7814 |
573.6 |
63.91 |
Amino acid frequency
Ala |
Cys |
Asp |
Glu |
Phe |
Gly |
His |
Ile |
Lys |
Leu |
|---|---|---|---|---|---|---|---|---|---|
6.972 ± 0.013 | 2.302 ± 0.014 |
4.829 ± 0.012 | 7.001 ± 0.018 |
3.754 ± 0.009 | 6.549 ± 0.018 |
2.579 ± 0.006 | 4.39 ± 0.011 |
5.623 ± 0.019 | 10.096 ± 0.02 |
Met |
Asn |
Gln |
Pro |
Arg |
Ser |
Thr |
Val |
Trp |
Tyr |
|---|---|---|---|---|---|---|---|---|---|
2.119 ± 0.005 | 3.604 ± 0.009 |
6.169 ± 0.021 | 4.707 ± 0.012 |
5.712 ± 0.012 | 8.196 ± 0.016 |
5.239 ± 0.009 | 6.18 ± 0.011 |
1.224 ± 0.005 | 2.689 ± 0.008 |
Most of the basic statistics you can see at this page can be downloaded from this CSV file
Proteome-pI is available under Creative Commons Attribution-NoDerivs license, for more details see here
| Reference: Kozlowski LP. Proteome-pI 2.0: Proteome Isoelectric Point Database Update. Nucleic Acids Res. 2021, doi: 10.1093/nar/gkab944 | Contact: Lukasz P. Kozlowski |
