Beijerinckiaceae bacterium
Taxonomy: cellular organisms; Bacteria; Proteobacteria; Alphaproteobacteria; Hyphomicrobiales; Beijerinckiaceae; unclassified Beijerinckiaceae
Average proteome isoelectric point is 7.18
Get precalculated fractions of proteins
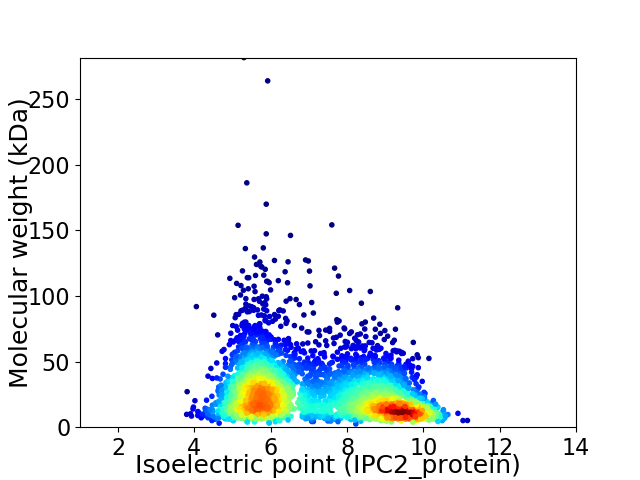
Virtual 2D-PAGE plot for 3557 proteins (isoelectric point calculated using IPC2_protein)
Get csv file with sequences according to given criteria:
* You can choose from 21 different methods for calculating isoelectric point
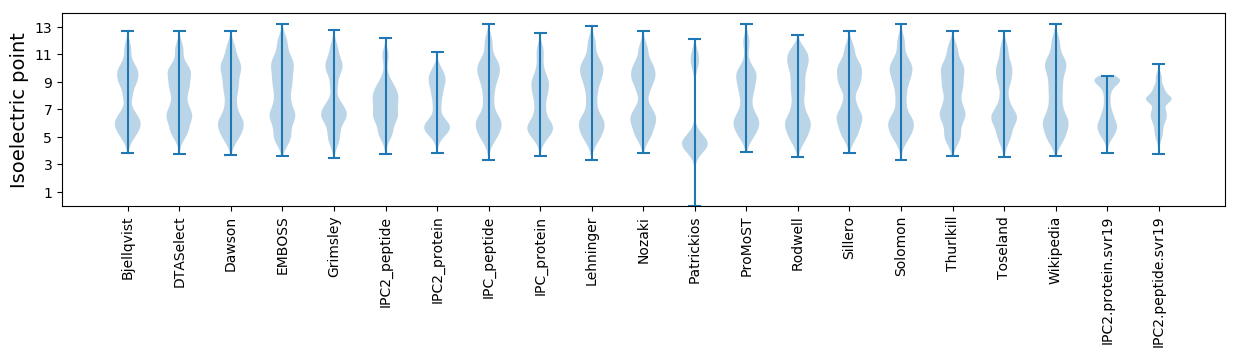
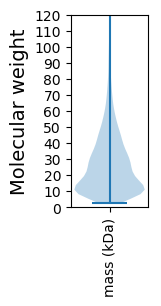
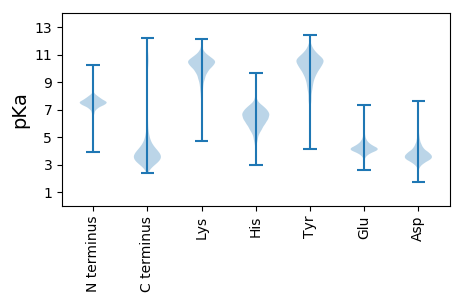
Summary statistics related to proteome-wise predictions
Protein with the lowest isoelectric point:
>tr|A0A2N8M7B3|A0A2N8M7B3_9RHIZ Uncharacterized protein (Fragment) OS=Beijerinckiaceae bacterium OX=1978229 GN=CR217_16820 PE=4 SV=1
MM1 pKa = 8.14DD2 pKa = 4.82SPAGAAPFVYY12 pKa = 9.14VTNSVSNTVSVIDD25 pKa = 3.68TATNTVVATIPAGVEE40 pKa = 3.72PAGVAVTPDD49 pKa = 3.1GKK51 pKa = 9.91HH52 pKa = 6.4AYY54 pKa = 9.24LANQGTLSVPGTTVLVIDD72 pKa = 3.9MTTNTVVATVGVGNFPSGVAITPGGTHH99 pKa = 6.09VYY101 pKa = 9.66VVNGFSNNVSVIDD114 pKa = 3.77TASNTVVGTPIPVGKK129 pKa = 10.04FPQSVALTPDD139 pKa = 3.38GTHH142 pKa = 5.92GFVVNEE148 pKa = 3.95FSNNVSVIDD157 pKa = 3.79TASNTMVGIPIPVGNFPIGVAITPNGTHH185 pKa = 7.07GYY187 pKa = 7.59VTNFDD192 pKa = 4.44DD193 pKa = 4.99NNVSVIDD200 pKa = 3.66TT201 pKa = 3.96
MM1 pKa = 8.14DD2 pKa = 4.82SPAGAAPFVYY12 pKa = 9.14VTNSVSNTVSVIDD25 pKa = 3.68TATNTVVATIPAGVEE40 pKa = 3.72PAGVAVTPDD49 pKa = 3.1GKK51 pKa = 9.91HH52 pKa = 6.4AYY54 pKa = 9.24LANQGTLSVPGTTVLVIDD72 pKa = 3.9MTTNTVVATVGVGNFPSGVAITPGGTHH99 pKa = 6.09VYY101 pKa = 9.66VVNGFSNNVSVIDD114 pKa = 3.77TASNTVVGTPIPVGKK129 pKa = 10.04FPQSVALTPDD139 pKa = 3.38GTHH142 pKa = 5.92GFVVNEE148 pKa = 3.95FSNNVSVIDD157 pKa = 3.79TASNTMVGIPIPVGNFPIGVAITPNGTHH185 pKa = 7.07GYY187 pKa = 7.59VTNFDD192 pKa = 4.44DD193 pKa = 4.99NNVSVIDD200 pKa = 3.66TT201 pKa = 3.96
Molecular weight: 20.2 kDa
Isoelectric point according different methods:
Protein with the highest isoelectric point:
>tr|A0A2N8MD69|A0A2N8MD69_9RHIZ Adenosine kinase OS=Beijerinckiaceae bacterium OX=1978229 GN=CR217_05575 PE=4 SV=1
MM1 pKa = 7.35KK2 pKa = 9.43RR3 pKa = 11.84TYY5 pKa = 10.27QPSKK9 pKa = 9.73LVRR12 pKa = 11.84KK13 pKa = 9.15RR14 pKa = 11.84RR15 pKa = 11.84HH16 pKa = 4.42GFRR19 pKa = 11.84ARR21 pKa = 11.84MASVGGRR28 pKa = 11.84KK29 pKa = 9.29VIAARR34 pKa = 11.84RR35 pKa = 11.84ARR37 pKa = 11.84GRR39 pKa = 11.84KK40 pKa = 9.03RR41 pKa = 11.84LSAA44 pKa = 4.03
MM1 pKa = 7.35KK2 pKa = 9.43RR3 pKa = 11.84TYY5 pKa = 10.27QPSKK9 pKa = 9.73LVRR12 pKa = 11.84KK13 pKa = 9.15RR14 pKa = 11.84RR15 pKa = 11.84HH16 pKa = 4.42GFRR19 pKa = 11.84ARR21 pKa = 11.84MASVGGRR28 pKa = 11.84KK29 pKa = 9.29VIAARR34 pKa = 11.84RR35 pKa = 11.84ARR37 pKa = 11.84GRR39 pKa = 11.84KK40 pKa = 9.03RR41 pKa = 11.84LSAA44 pKa = 4.03
Molecular weight: 5.12 kDa
Isoelectric point according different methods:
Peptides (in silico digests for buttom-up proteomics)
Below you can find in silico digests of the whole proteome with Trypsin, Chymotrypsin, Trypsin+LysC, LysN, ArgC proteases suitable for different mass spec machines.| Try ESI |
 |
|---|
| ChTry ESI |
 |
|---|
| ArgC ESI |
 |
|---|
| LysN ESI |
 |
|---|
| TryLysC ESI |
 |
|---|
| Try MALDI |
 |
|---|
| ChTry MALDI |
 |
|---|
| ArgC MALDI |
 |
|---|
| LysN MALDI |
 |
|---|
| TryLysC MALDI |
 |
|---|
| Try LTQ |
 |
|---|
| ChTry LTQ |
 |
|---|
| ArgC LTQ |
 |
|---|
| LysN LTQ |
 |
|---|
| TryLysC LTQ |
 |
|---|
| Try MSlow |
 |
|---|
| ChTry MSlow |
 |
|---|
| ArgC MSlow |
 |
|---|
| LysN MSlow |
 |
|---|
| TryLysC MSlow |
 |
|---|
| Try MShigh |
 |
|---|
| ChTry MShigh |
 |
|---|
| ArgC MShigh |
 |
|---|
| LysN MShigh |
 |
|---|
| TryLysC MShigh |
 |
|---|
General Statistics
Number of major isoforms |
Number of additional isoforms |
Number of all proteins |
Number of amino acids |
Min. Seq. Length |
Max. Seq. Length |
Avg. Seq. Length |
Avg. Mol. Weight |
|---|---|---|---|---|---|---|---|
0 |
927747 |
26 |
2626 |
260.8 |
28.35 |
Amino acid frequency
Ala |
Cys |
Asp |
Glu |
Phe |
Gly |
His |
Ile |
Lys |
Leu |
|---|---|---|---|---|---|---|---|---|---|
12.837 ± 0.058 | 1.037 ± 0.016 |
5.26 ± 0.03 | 5.55 ± 0.049 |
3.981 ± 0.034 | 8.393 ± 0.042 |
2.163 ± 0.02 | 5.309 ± 0.029 |
3.728 ± 0.038 | 10.025 ± 0.051 |
Met |
Asn |
Gln |
Pro |
Arg |
Ser |
Thr |
Val |
Trp |
Tyr |
|---|---|---|---|---|---|---|---|---|---|
2.177 ± 0.016 | 2.766 ± 0.026 |
5.515 ± 0.04 | 2.95 ± 0.026 |
7.472 ± 0.049 | 5.479 ± 0.028 |
5.0 ± 0.027 | 6.858 ± 0.036 |
1.319 ± 0.02 | 2.18 ± 0.023 |
Most of the basic statistics you can see at this page can be downloaded from this CSV file
Proteome-pI is available under Creative Commons Attribution-NoDerivs license, for more details see here
| Reference: Kozlowski LP. Proteome-pI 2.0: Proteome Isoelectric Point Database Update. Nucleic Acids Res. 2021, doi: 10.1093/nar/gkab944 | Contact: Lukasz P. Kozlowski |
