Penicillium nordicum
Taxonomy: cellular organisms; Eukaryota; Opisthokonta; Fungi; Dikarya; Ascomycota; saccharomyceta; Pezizomycotina; leotiomyceta; Eurotiomycetes; Eurotiomycetidae; Eurotiales; Aspergillaceae; Penicillium
Average proteome isoelectric point is 6.6
Get precalculated fractions of proteins
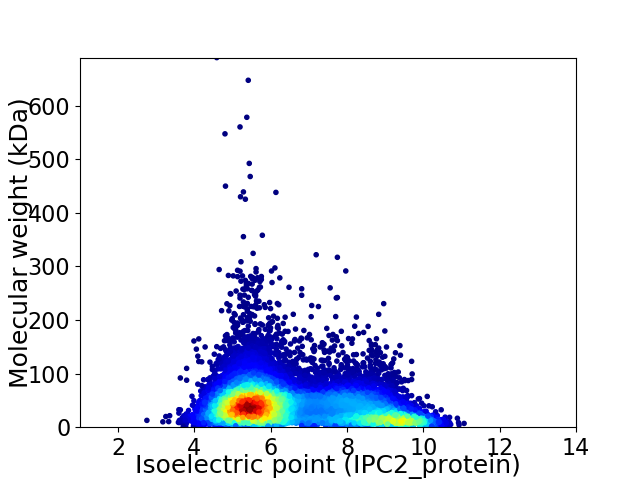
Virtual 2D-PAGE plot for 12938 proteins (isoelectric point calculated using IPC2_protein)
Get csv file with sequences according to given criteria:
* You can choose from 21 different methods for calculating isoelectric point
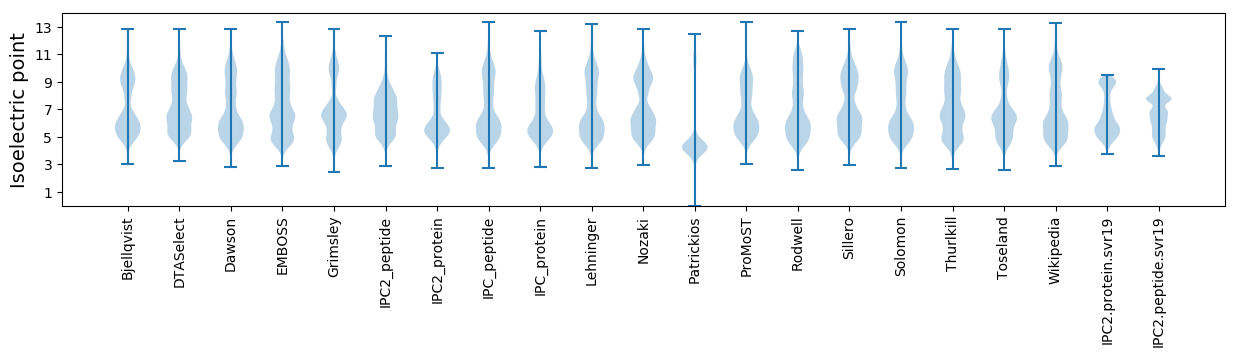
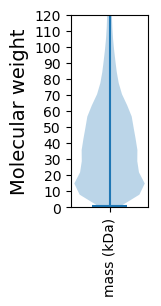
Summary statistics related to proteome-wise predictions
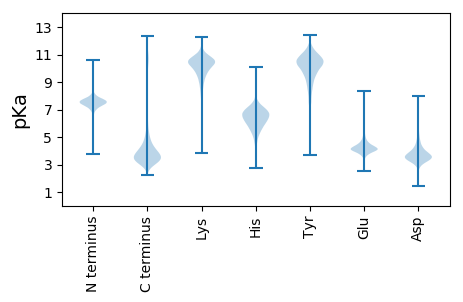
Protein with the lowest isoelectric point:
>tr|A0A0M8P025|A0A0M8P025_9EURO Uncharacterized protein OS=Penicillium nordicum OX=229535 GN=ACN38_g8358 PE=4 SV=1
MM1 pKa = 7.13TFNVFLTGLLAATAAALPHH20 pKa = 5.95SARR23 pKa = 11.84SGEE26 pKa = 4.15SGAVEE31 pKa = 4.25TRR33 pKa = 11.84DD34 pKa = 3.57SSSGGLQIVNNMDD47 pKa = 3.23TDD49 pKa = 4.02VYY51 pKa = 11.16LWTTSSDD58 pKa = 3.31AGSMKK63 pKa = 10.11TLSTGGDD70 pKa = 3.79YY71 pKa = 11.37SEE73 pKa = 5.77DD74 pKa = 3.05WATNSNGGGISIKK87 pKa = 10.18MSTSEE92 pKa = 4.29SKK94 pKa = 11.07DD95 pKa = 3.2SVLQFEE101 pKa = 4.64YY102 pKa = 10.25TQDD105 pKa = 3.43GDD107 pKa = 4.04TLFWDD112 pKa = 4.11MSSIDD117 pKa = 4.43LDD119 pKa = 3.86STSEE123 pKa = 4.0FVKK126 pKa = 10.8SGFTVVPSDD135 pKa = 4.0SSCKK139 pKa = 10.14SVSCAAGDD147 pKa = 4.17AEE149 pKa = 5.81CKK151 pKa = 10.0DD152 pKa = 4.38AYY154 pKa = 10.57QLPDD158 pKa = 3.43DD159 pKa = 4.56VNTYY163 pKa = 10.49SCSLSSGFTLTLGG176 pKa = 3.59
MM1 pKa = 7.13TFNVFLTGLLAATAAALPHH20 pKa = 5.95SARR23 pKa = 11.84SGEE26 pKa = 4.15SGAVEE31 pKa = 4.25TRR33 pKa = 11.84DD34 pKa = 3.57SSSGGLQIVNNMDD47 pKa = 3.23TDD49 pKa = 4.02VYY51 pKa = 11.16LWTTSSDD58 pKa = 3.31AGSMKK63 pKa = 10.11TLSTGGDD70 pKa = 3.79YY71 pKa = 11.37SEE73 pKa = 5.77DD74 pKa = 3.05WATNSNGGGISIKK87 pKa = 10.18MSTSEE92 pKa = 4.29SKK94 pKa = 11.07DD95 pKa = 3.2SVLQFEE101 pKa = 4.64YY102 pKa = 10.25TQDD105 pKa = 3.43GDD107 pKa = 4.04TLFWDD112 pKa = 4.11MSSIDD117 pKa = 4.43LDD119 pKa = 3.86STSEE123 pKa = 4.0FVKK126 pKa = 10.8SGFTVVPSDD135 pKa = 4.0SSCKK139 pKa = 10.14SVSCAAGDD147 pKa = 4.17AEE149 pKa = 5.81CKK151 pKa = 10.0DD152 pKa = 4.38AYY154 pKa = 10.57QLPDD158 pKa = 3.43DD159 pKa = 4.56VNTYY163 pKa = 10.49SCSLSSGFTLTLGG176 pKa = 3.59
Molecular weight: 18.44 kDa
Isoelectric point according different methods:
Protein with the highest isoelectric point:
>tr|A0A0M8P0P1|A0A0M8P0P1_9EURO Non-specific serine/threonine protein kinase OS=Penicillium nordicum OX=229535 GN=ACN38_g10971 PE=4 SV=1
MM1 pKa = 7.49PVSPSRR7 pKa = 11.84TRR9 pKa = 11.84PSTAGRR15 pKa = 11.84TMSTTTSVPLPRR27 pKa = 11.84ARR29 pKa = 11.84TSVPASSSTTPSALFAPRR47 pKa = 11.84PGPIVGTVSARR58 pKa = 11.84LATSLSTLTSRR69 pKa = 5.13
MM1 pKa = 7.49PVSPSRR7 pKa = 11.84TRR9 pKa = 11.84PSTAGRR15 pKa = 11.84TMSTTTSVPLPRR27 pKa = 11.84ARR29 pKa = 11.84TSVPASSSTTPSALFAPRR47 pKa = 11.84PGPIVGTVSARR58 pKa = 11.84LATSLSTLTSRR69 pKa = 5.13
Molecular weight: 7.03 kDa
Isoelectric point according different methods:
Peptides (in silico digests for buttom-up proteomics)
Below you can find in silico digests of the whole proteome with Trypsin, Chymotrypsin, Trypsin+LysC, LysN, ArgC proteases suitable for different mass spec machines.| Try ESI |
 |
|---|
| ChTry ESI |
 |
|---|
| ArgC ESI |
 |
|---|
| LysN ESI |
 |
|---|
| TryLysC ESI |
 |
|---|
| Try MALDI |
 |
|---|
| ChTry MALDI |
 |
|---|
| ArgC MALDI |
 |
|---|
| LysN MALDI |
 |
|---|
| TryLysC MALDI |
 |
|---|
| Try LTQ |
 |
|---|
| ChTry LTQ |
 |
|---|
| ArgC LTQ |
 |
|---|
| LysN LTQ |
 |
|---|
| TryLysC LTQ |
 |
|---|
| Try MSlow |
 |
|---|
| ChTry MSlow |
 |
|---|
| ArgC MSlow |
 |
|---|
| LysN MSlow |
 |
|---|
| TryLysC MSlow |
 |
|---|
| Try MShigh |
 |
|---|
| ChTry MShigh |
 |
|---|
| ArgC MShigh |
 |
|---|
| LysN MShigh |
 |
|---|
| TryLysC MShigh |
 |
|---|
General Statistics
Number of major isoforms |
Number of additional isoforms |
Number of all proteins |
Number of amino acids |
Min. Seq. Length |
Max. Seq. Length |
Avg. Seq. Length |
Avg. Mol. Weight |
|---|---|---|---|---|---|---|---|
0 |
5328948 |
8 |
6304 |
411.9 |
45.67 |
Amino acid frequency
Ala |
Cys |
Asp |
Glu |
Phe |
Gly |
His |
Ile |
Lys |
Leu |
|---|---|---|---|---|---|---|---|---|---|
8.189 ± 0.021 | 1.283 ± 0.009 |
5.596 ± 0.018 | 6.023 ± 0.026 |
3.872 ± 0.015 | 6.713 ± 0.02 |
2.473 ± 0.01 | 5.168 ± 0.018 |
4.649 ± 0.02 | 9.037 ± 0.025 |
Met |
Asn |
Gln |
Pro |
Arg |
Ser |
Thr |
Val |
Trp |
Tyr |
|---|---|---|---|---|---|---|---|---|---|
2.272 ± 0.008 | 3.762 ± 0.012 |
6.072 ± 0.025 | 4.048 ± 0.018 |
5.961 ± 0.02 | 8.449 ± 0.024 |
6.021 ± 0.014 | 6.125 ± 0.016 |
1.469 ± 0.009 | 2.814 ± 0.012 |
Most of the basic statistics you can see at this page can be downloaded from this CSV file
Proteome-pI is available under Creative Commons Attribution-NoDerivs license, for more details see here
| Reference: Kozlowski LP. Proteome-pI 2.0: Proteome Isoelectric Point Database Update. Nucleic Acids Res. 2021, doi: 10.1093/nar/gkab944 | Contact: Lukasz P. Kozlowski |
