Streptomyces paucisporeus
Taxonomy: cellular organisms; Bacteria; Terrabacteria group; Actinobacteria; Actinomycetia; Streptomycetales; Streptomycetaceae; Streptomyces
Average proteome isoelectric point is 6.49
Get precalculated fractions of proteins
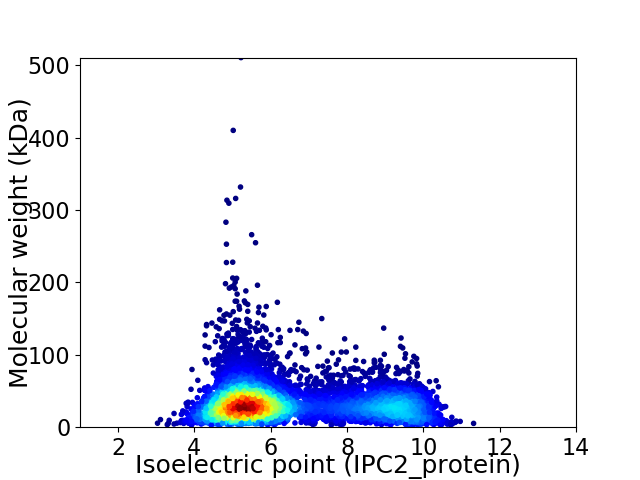
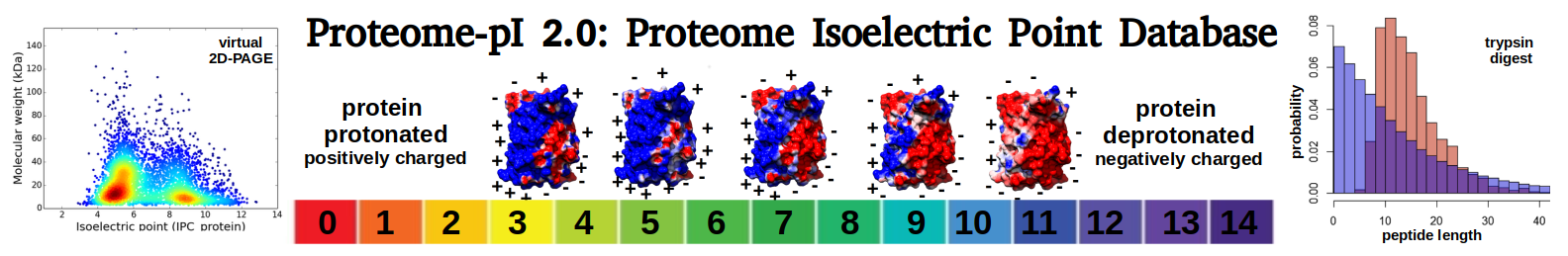
Virtual 2D-PAGE plot for 7034 proteins (isoelectric point calculated using IPC2_protein)
Get csv file with sequences according to given criteria:
* You can choose from 21 different methods for calculating isoelectric point
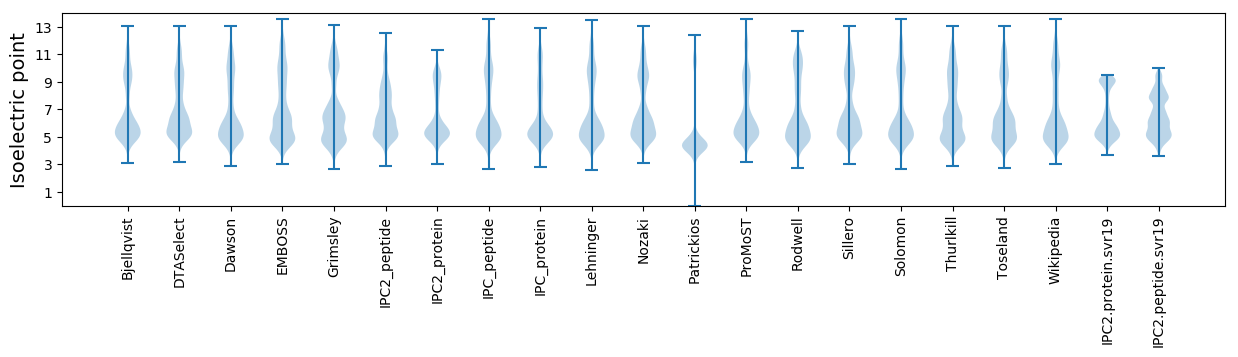
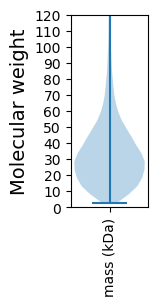
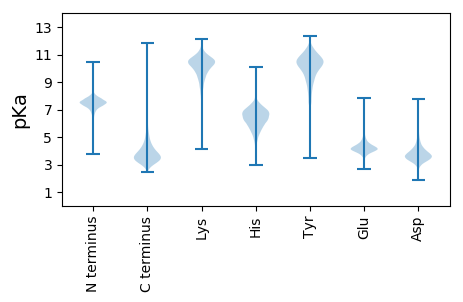
Summary statistics related to proteome-wise predictions
Protein with the lowest isoelectric point:
>tr|A0A1M6WSL4|A0A1M6WSL4_9ACTN Energy-dependent translational throttle protein EttA OS=Streptomyces paucisporeus OX=310782 GN=ettA PE=3 SV=1
MM1 pKa = 7.35ARR3 pKa = 11.84WDD5 pKa = 3.92AEE7 pKa = 4.15NQRR10 pKa = 11.84WAQDD14 pKa = 3.94PPPGAAGGPGPASVPRR30 pKa = 11.84TPGGNSRR37 pKa = 11.84VLIVALAVVLVCAAAGIGLWAAQTDD62 pKa = 3.92SGGGGGSSPAAGAPGFGDD80 pKa = 3.55GTSAYY85 pKa = 10.61SPDD88 pKa = 3.83PATAGDD94 pKa = 4.41SDD96 pKa = 4.24TTDD99 pKa = 3.37LPTTEE104 pKa = 5.83DD105 pKa = 3.67PTTDD109 pKa = 3.58FPTTEE114 pKa = 4.58DD115 pKa = 4.15PSTEE119 pKa = 4.07PATTGPPPGFAHH131 pKa = 6.32LQDD134 pKa = 3.7VAGFTLDD141 pKa = 4.84VPEE144 pKa = 4.07SWQRR148 pKa = 11.84SSGGASVYY156 pKa = 10.22YY157 pKa = 9.9QSQDD161 pKa = 3.31QQGLIQVFSLSSPQTTPYY179 pKa = 10.49EE180 pKa = 4.19SLKK183 pKa = 10.07ATEE186 pKa = 4.64ATVSTNNGYY195 pKa = 9.19QLLGLRR201 pKa = 11.84YY202 pKa = 9.72IIDD205 pKa = 4.16DD206 pKa = 3.92QASDD210 pKa = 3.79AADD213 pKa = 3.47LEE215 pKa = 4.55YY216 pKa = 10.39TYY218 pKa = 11.42VRR220 pKa = 11.84DD221 pKa = 4.19DD222 pKa = 3.25GGTRR226 pKa = 11.84HH227 pKa = 5.29VVDD230 pKa = 4.5RR231 pKa = 11.84AFTGPDD237 pKa = 3.2GVQYY241 pKa = 11.25ALLVAGPDD249 pKa = 4.09SDD251 pKa = 3.05WSSYY255 pKa = 9.69QDD257 pKa = 3.17VFRR260 pKa = 11.84VLRR263 pKa = 11.84ASFCPDD269 pKa = 3.2NYY271 pKa = 9.73CTQQ274 pKa = 3.17
MM1 pKa = 7.35ARR3 pKa = 11.84WDD5 pKa = 3.92AEE7 pKa = 4.15NQRR10 pKa = 11.84WAQDD14 pKa = 3.94PPPGAAGGPGPASVPRR30 pKa = 11.84TPGGNSRR37 pKa = 11.84VLIVALAVVLVCAAAGIGLWAAQTDD62 pKa = 3.92SGGGGGSSPAAGAPGFGDD80 pKa = 3.55GTSAYY85 pKa = 10.61SPDD88 pKa = 3.83PATAGDD94 pKa = 4.41SDD96 pKa = 4.24TTDD99 pKa = 3.37LPTTEE104 pKa = 5.83DD105 pKa = 3.67PTTDD109 pKa = 3.58FPTTEE114 pKa = 4.58DD115 pKa = 4.15PSTEE119 pKa = 4.07PATTGPPPGFAHH131 pKa = 6.32LQDD134 pKa = 3.7VAGFTLDD141 pKa = 4.84VPEE144 pKa = 4.07SWQRR148 pKa = 11.84SSGGASVYY156 pKa = 10.22YY157 pKa = 9.9QSQDD161 pKa = 3.31QQGLIQVFSLSSPQTTPYY179 pKa = 10.49EE180 pKa = 4.19SLKK183 pKa = 10.07ATEE186 pKa = 4.64ATVSTNNGYY195 pKa = 9.19QLLGLRR201 pKa = 11.84YY202 pKa = 9.72IIDD205 pKa = 4.16DD206 pKa = 3.92QASDD210 pKa = 3.79AADD213 pKa = 3.47LEE215 pKa = 4.55YY216 pKa = 10.39TYY218 pKa = 11.42VRR220 pKa = 11.84DD221 pKa = 4.19DD222 pKa = 3.25GGTRR226 pKa = 11.84HH227 pKa = 5.29VVDD230 pKa = 4.5RR231 pKa = 11.84AFTGPDD237 pKa = 3.2GVQYY241 pKa = 11.25ALLVAGPDD249 pKa = 4.09SDD251 pKa = 3.05WSSYY255 pKa = 9.69QDD257 pKa = 3.17VFRR260 pKa = 11.84VLRR263 pKa = 11.84ASFCPDD269 pKa = 3.2NYY271 pKa = 9.73CTQQ274 pKa = 3.17
Molecular weight: 28.49 kDa
Isoelectric point according different methods:
Protein with the highest isoelectric point:
>tr|A0A1M7CFI9|A0A1M7CFI9_9ACTN Uncharacterized protein OS=Streptomyces paucisporeus OX=310782 GN=SAMN05216499_105224 PE=4 SV=1
MM1 pKa = 7.69SKK3 pKa = 9.0RR4 pKa = 11.84TFQPNNRR11 pKa = 11.84RR12 pKa = 11.84RR13 pKa = 11.84AKK15 pKa = 8.7THH17 pKa = 5.15GFRR20 pKa = 11.84LRR22 pKa = 11.84MRR24 pKa = 11.84TRR26 pKa = 11.84AGRR29 pKa = 11.84AILATRR35 pKa = 11.84RR36 pKa = 11.84GKK38 pKa = 10.48GRR40 pKa = 11.84ARR42 pKa = 11.84LSAA45 pKa = 3.91
MM1 pKa = 7.69SKK3 pKa = 9.0RR4 pKa = 11.84TFQPNNRR11 pKa = 11.84RR12 pKa = 11.84RR13 pKa = 11.84AKK15 pKa = 8.7THH17 pKa = 5.15GFRR20 pKa = 11.84LRR22 pKa = 11.84MRR24 pKa = 11.84TRR26 pKa = 11.84AGRR29 pKa = 11.84AILATRR35 pKa = 11.84RR36 pKa = 11.84GKK38 pKa = 10.48GRR40 pKa = 11.84ARR42 pKa = 11.84LSAA45 pKa = 3.91
Molecular weight: 5.27 kDa
Isoelectric point according different methods:
Peptides (in silico digests for buttom-up proteomics)
Below you can find in silico digests of the whole proteome with Trypsin, Chymotrypsin, Trypsin+LysC, LysN, ArgC proteases suitable for different mass spec machines.| Try ESI |
 |
|---|
| ChTry ESI |
 |
|---|
| ArgC ESI |
 |
|---|
| LysN ESI |
 |
|---|
| TryLysC ESI |
 |
|---|
| Try MALDI |
 |
|---|
| ChTry MALDI |
 |
|---|
| ArgC MALDI |
 |
|---|
| LysN MALDI |
 |
|---|
| TryLysC MALDI |
 |
|---|
| Try LTQ |
 |
|---|
| ChTry LTQ |
 |
|---|
| ArgC LTQ |
 |
|---|
| LysN LTQ |
 |
|---|
| TryLysC LTQ |
 |
|---|
| Try MSlow |
 |
|---|
| ChTry MSlow |
 |
|---|
| ArgC MSlow |
 |
|---|
| LysN MSlow |
 |
|---|
| TryLysC MSlow |
 |
|---|
| Try MShigh |
 |
|---|
| ChTry MShigh |
 |
|---|
| ArgC MShigh |
 |
|---|
| LysN MShigh |
 |
|---|
| TryLysC MShigh |
 |
|---|
General Statistics
Number of major isoforms |
Number of additional isoforms |
Number of all proteins |
Number of amino acids |
Min. Seq. Length |
Max. Seq. Length |
Avg. Seq. Length |
Avg. Mol. Weight |
|---|---|---|---|---|---|---|---|
0 |
2374895 |
27 |
4814 |
337.6 |
35.88 |
Amino acid frequency
Ala |
Cys |
Asp |
Glu |
Phe |
Gly |
His |
Ile |
Lys |
Leu |
|---|---|---|---|---|---|---|---|---|---|
14.412 ± 0.038 | 0.784 ± 0.008 |
6.032 ± 0.024 | 4.912 ± 0.034 |
2.662 ± 0.016 | 9.744 ± 0.024 |
2.323 ± 0.014 | 3.008 ± 0.019 |
1.862 ± 0.021 | 10.172 ± 0.037 |
Met |
Asn |
Gln |
Pro |
Arg |
Ser |
Thr |
Val |
Trp |
Tyr |
|---|---|---|---|---|---|---|---|---|---|
1.657 ± 0.013 | 1.801 ± 0.019 |
6.331 ± 0.026 | 2.903 ± 0.019 |
7.867 ± 0.038 | 5.179 ± 0.027 |
6.367 ± 0.032 | 8.329 ± 0.027 |
1.524 ± 0.012 | 2.129 ± 0.015 |
Most of the basic statistics you can see at this page can be downloaded from this CSV file
Proteome-pI is available under Creative Commons Attribution-NoDerivs license, for more details see here
| Reference: Kozlowski LP. Proteome-pI 2.0: Proteome Isoelectric Point Database Update. Nucleic Acids Res. 2021, doi: 10.1093/nar/gkab944 | Contact: Lukasz P. Kozlowski |
