Extensimonas vulgaris
Taxonomy: cellular organisms; Bacteria; Proteobacteria; Betaproteobacteria; Burkholderiales; Comamonadaceae; Extensimonas
Average proteome isoelectric point is 6.95
Get precalculated fractions of proteins
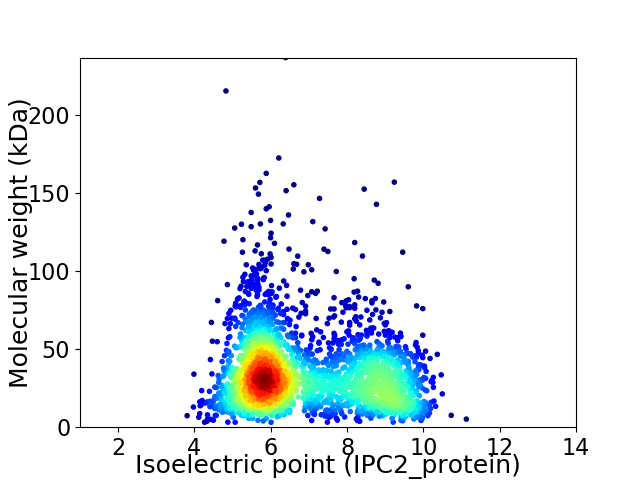
Virtual 2D-PAGE plot for 2789 proteins (isoelectric point calculated using IPC2_protein)
Get csv file with sequences according to given criteria:
* You can choose from 21 different methods for calculating isoelectric point
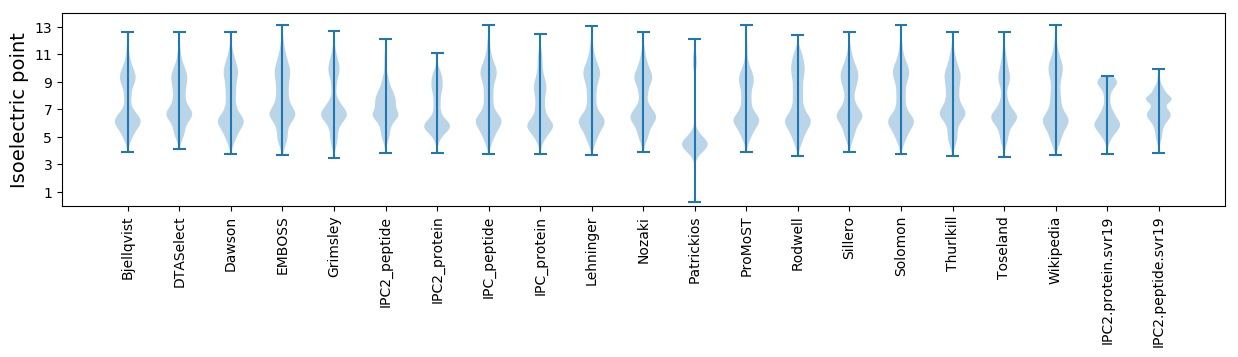
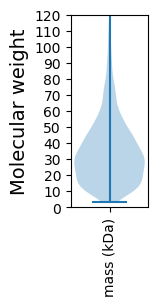
Summary statistics related to proteome-wise predictions
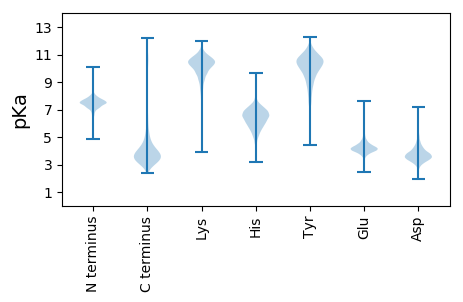
Protein with the lowest isoelectric point:
>tr|A0A369AFR7|A0A369AFR7_9BURK Type IV secretory system conjugative DNA transfer VirD4/TraG family protein (Fragment) OS=Extensimonas vulgaris OX=1031594 GN=DFR45_11724 PE=4 SV=1
MM1 pKa = 7.25SAVAEE6 pKa = 4.48NISTEE11 pKa = 3.97MPAPINFTDD20 pKa = 3.42SAAAKK25 pKa = 9.27VAEE28 pKa = 5.01LIAEE32 pKa = 4.35EE33 pKa = 4.62GNPDD37 pKa = 2.99LKK39 pKa = 11.13LRR41 pKa = 11.84VFVQGGGCSGFQYY54 pKa = 10.9GFTFDD59 pKa = 3.8EE60 pKa = 4.88VVNEE64 pKa = 4.95DD65 pKa = 3.69DD66 pKa = 3.27TTMVKK71 pKa = 10.58NGVTLLIDD79 pKa = 3.64AMSYY83 pKa = 10.32QYY85 pKa = 11.5LVGAEE90 pKa = 3.79IDD92 pKa = 3.91YY93 pKa = 11.36KK94 pKa = 11.19EE95 pKa = 4.67DD96 pKa = 3.23LQGAQFVIRR105 pKa = 11.84NPNATSTCGCGSSFSVV121 pKa = 3.54
MM1 pKa = 7.25SAVAEE6 pKa = 4.48NISTEE11 pKa = 3.97MPAPINFTDD20 pKa = 3.42SAAAKK25 pKa = 9.27VAEE28 pKa = 5.01LIAEE32 pKa = 4.35EE33 pKa = 4.62GNPDD37 pKa = 2.99LKK39 pKa = 11.13LRR41 pKa = 11.84VFVQGGGCSGFQYY54 pKa = 10.9GFTFDD59 pKa = 3.8EE60 pKa = 4.88VVNEE64 pKa = 4.95DD65 pKa = 3.69DD66 pKa = 3.27TTMVKK71 pKa = 10.58NGVTLLIDD79 pKa = 3.64AMSYY83 pKa = 10.32QYY85 pKa = 11.5LVGAEE90 pKa = 3.79IDD92 pKa = 3.91YY93 pKa = 11.36KK94 pKa = 11.19EE95 pKa = 4.67DD96 pKa = 3.23LQGAQFVIRR105 pKa = 11.84NPNATSTCGCGSSFSVV121 pKa = 3.54
Molecular weight: 12.91 kDa
Isoelectric point according different methods:
Protein with the highest isoelectric point:
>tr|A0A369AIX7|A0A369AIX7_9BURK Uncharacterized protein OS=Extensimonas vulgaris OX=1031594 GN=DFR45_109109 PE=4 SV=1
MM1 pKa = 7.35KK2 pKa = 9.43RR3 pKa = 11.84TYY5 pKa = 10.25QPSKK9 pKa = 7.79TRR11 pKa = 11.84RR12 pKa = 11.84ARR14 pKa = 11.84THH16 pKa = 5.79GFLVRR21 pKa = 11.84MKK23 pKa = 9.1TRR25 pKa = 11.84SGRR28 pKa = 11.84AIINARR34 pKa = 11.84RR35 pKa = 11.84AKK37 pKa = 9.62GRR39 pKa = 11.84KK40 pKa = 8.75RR41 pKa = 11.84LAVV44 pKa = 3.41
MM1 pKa = 7.35KK2 pKa = 9.43RR3 pKa = 11.84TYY5 pKa = 10.25QPSKK9 pKa = 7.79TRR11 pKa = 11.84RR12 pKa = 11.84ARR14 pKa = 11.84THH16 pKa = 5.79GFLVRR21 pKa = 11.84MKK23 pKa = 9.1TRR25 pKa = 11.84SGRR28 pKa = 11.84AIINARR34 pKa = 11.84RR35 pKa = 11.84AKK37 pKa = 9.62GRR39 pKa = 11.84KK40 pKa = 8.75RR41 pKa = 11.84LAVV44 pKa = 3.41
Molecular weight: 5.18 kDa
Isoelectric point according different methods:
Peptides (in silico digests for buttom-up proteomics)
Below you can find in silico digests of the whole proteome with Trypsin, Chymotrypsin, Trypsin+LysC, LysN, ArgC proteases suitable for different mass spec machines.| Try ESI |
 |
|---|
| ChTry ESI |
 |
|---|
| ArgC ESI |
 |
|---|
| LysN ESI |
 |
|---|
| TryLysC ESI |
 |
|---|
| Try MALDI |
 |
|---|
| ChTry MALDI |
 |
|---|
| ArgC MALDI |
 |
|---|
| LysN MALDI |
 |
|---|
| TryLysC MALDI |
 |
|---|
| Try LTQ |
 |
|---|
| ChTry LTQ |
 |
|---|
| ArgC LTQ |
 |
|---|
| LysN LTQ |
 |
|---|
| TryLysC LTQ |
 |
|---|
| Try MSlow |
 |
|---|
| ChTry MSlow |
 |
|---|
| ArgC MSlow |
 |
|---|
| LysN MSlow |
 |
|---|
| TryLysC MSlow |
 |
|---|
| Try MShigh |
 |
|---|
| ChTry MShigh |
 |
|---|
| ArgC MShigh |
 |
|---|
| LysN MShigh |
 |
|---|
| TryLysC MShigh |
 |
|---|
General Statistics
Number of major isoforms |
Number of additional isoforms |
Number of all proteins |
Number of amino acids |
Min. Seq. Length |
Max. Seq. Length |
Avg. Seq. Length |
Avg. Mol. Weight |
|---|---|---|---|---|---|---|---|
0 |
930153 |
28 |
2184 |
333.5 |
36.2 |
Amino acid frequency
Ala |
Cys |
Asp |
Glu |
Phe |
Gly |
His |
Ile |
Lys |
Leu |
|---|---|---|---|---|---|---|---|---|---|
13.724 ± 0.074 | 0.982 ± 0.015 |
4.891 ± 0.032 | 5.391 ± 0.043 |
3.388 ± 0.028 | 8.087 ± 0.046 |
2.385 ± 0.022 | 4.237 ± 0.038 |
3.122 ± 0.042 | 11.015 ± 0.069 |
Met |
Asn |
Gln |
Pro |
Arg |
Ser |
Thr |
Val |
Trp |
Tyr |
|---|---|---|---|---|---|---|---|---|---|
2.342 ± 0.021 | 2.373 ± 0.031 |
5.545 ± 0.034 | 4.587 ± 0.034 |
7.022 ± 0.041 | 4.821 ± 0.031 |
4.967 ± 0.032 | 7.366 ± 0.041 |
1.501 ± 0.024 | 2.253 ± 0.022 |
Most of the basic statistics you can see at this page can be downloaded from this CSV file
Proteome-pI is available under Creative Commons Attribution-NoDerivs license, for more details see here
| Reference: Kozlowski LP. Proteome-pI 2.0: Proteome Isoelectric Point Database Update. Nucleic Acids Res. 2021, doi: 10.1093/nar/gkab944 | Contact: Lukasz P. Kozlowski |
