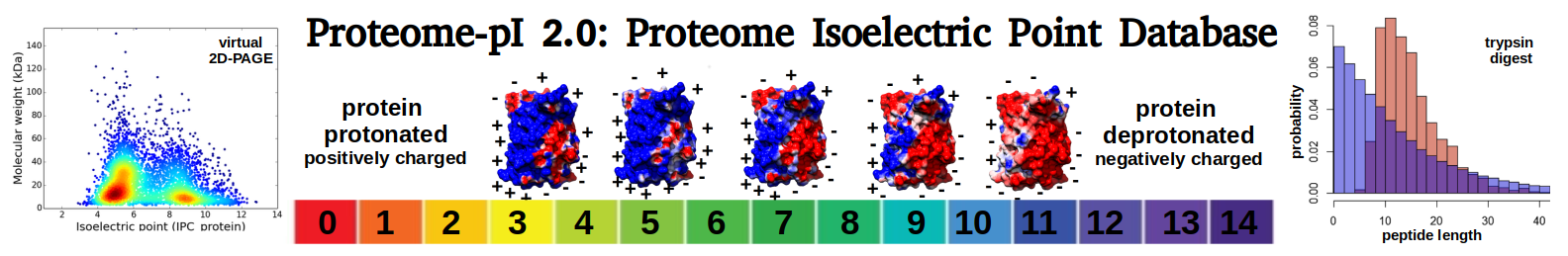
Streptococcus infantis SK970
Taxonomy: cellular organisms; Bacteria; Terrabacteria group; Firmicutes; Bacilli; Lactobacillales; Streptococcaceae; Streptococcus; Streptococcus infantis
Average proteome isoelectric point is 6.19
Get precalculated fractions of proteins
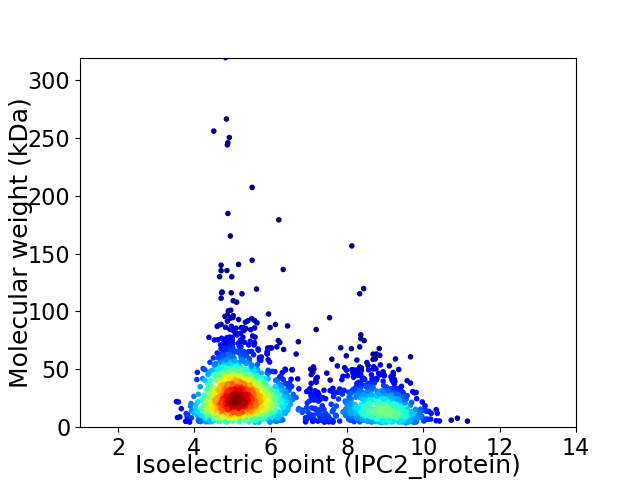
Virtual 2D-PAGE plot for 2144 proteins (isoelectric point calculated using IPC2_protein)
Get csv file with sequences according to given criteria:
* You can choose from 21 different methods for calculating isoelectric point
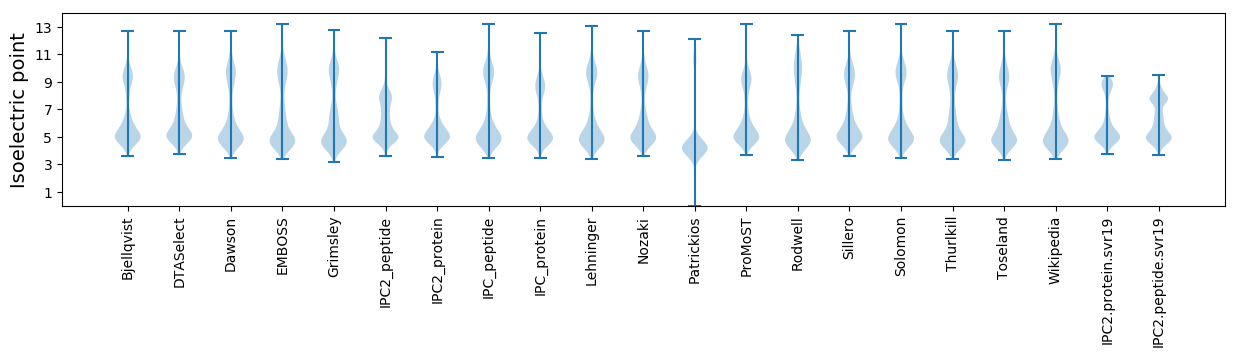
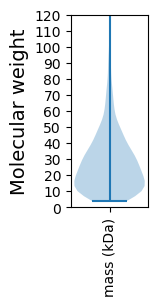
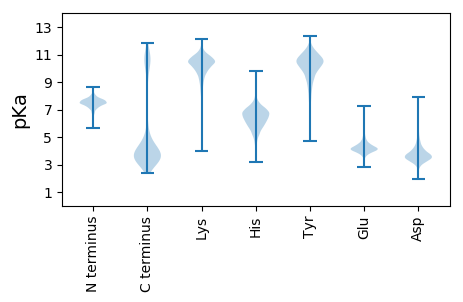
Summary statistics related to proteome-wise predictions
Protein with the lowest isoelectric point:
>tr|F9PUJ1|F9PUJ1_9STRE ABC transporter permease protein OS=Streptococcus infantis SK970 OX=1035189 GN=HMPREF9954_0895 PE=3 SV=1
MM1 pKa = 7.28KK2 pKa = 9.87RR3 pKa = 11.84DD4 pKa = 3.37EE5 pKa = 5.08FGFILPNLYY14 pKa = 10.4YY15 pKa = 10.62QFLTEE20 pKa = 4.68LKK22 pKa = 10.21EE23 pKa = 3.69IDD25 pKa = 3.98PYY27 pKa = 11.28EE28 pKa = 5.54IGDD31 pKa = 4.33SGICLYY37 pKa = 10.78AQEE40 pKa = 5.16DD41 pKa = 3.74LQEE44 pKa = 4.4RR45 pKa = 11.84NEE47 pKa = 4.19TYY49 pKa = 10.3QIEE52 pKa = 4.57EE53 pKa = 4.14VEE55 pKa = 4.17PDD57 pKa = 3.28YY58 pKa = 11.59FMIGQEE64 pKa = 3.56GDD66 pKa = 2.95LAYY69 pKa = 10.23FIKK72 pKa = 10.68KK73 pKa = 10.41NSDD76 pKa = 3.05DD77 pKa = 4.51CIYY80 pKa = 10.94EE81 pKa = 4.03NDD83 pKa = 3.68LGALGSLEE91 pKa = 4.12MQKK94 pKa = 10.32VAANVYY100 pKa = 10.61DD101 pKa = 4.89FIEE104 pKa = 4.4KK105 pKa = 9.97VLKK108 pKa = 10.81EE109 pKa = 4.04EE110 pKa = 4.26LL111 pKa = 3.56
MM1 pKa = 7.28KK2 pKa = 9.87RR3 pKa = 11.84DD4 pKa = 3.37EE5 pKa = 5.08FGFILPNLYY14 pKa = 10.4YY15 pKa = 10.62QFLTEE20 pKa = 4.68LKK22 pKa = 10.21EE23 pKa = 3.69IDD25 pKa = 3.98PYY27 pKa = 11.28EE28 pKa = 5.54IGDD31 pKa = 4.33SGICLYY37 pKa = 10.78AQEE40 pKa = 5.16DD41 pKa = 3.74LQEE44 pKa = 4.4RR45 pKa = 11.84NEE47 pKa = 4.19TYY49 pKa = 10.3QIEE52 pKa = 4.57EE53 pKa = 4.14VEE55 pKa = 4.17PDD57 pKa = 3.28YY58 pKa = 11.59FMIGQEE64 pKa = 3.56GDD66 pKa = 2.95LAYY69 pKa = 10.23FIKK72 pKa = 10.68KK73 pKa = 10.41NSDD76 pKa = 3.05DD77 pKa = 4.51CIYY80 pKa = 10.94EE81 pKa = 4.03NDD83 pKa = 3.68LGALGSLEE91 pKa = 4.12MQKK94 pKa = 10.32VAANVYY100 pKa = 10.61DD101 pKa = 4.89FIEE104 pKa = 4.4KK105 pKa = 9.97VLKK108 pKa = 10.81EE109 pKa = 4.04EE110 pKa = 4.26LL111 pKa = 3.56
Molecular weight: 13.02 kDa
Isoelectric point according different methods:
Protein with the highest isoelectric point:
>tr|F9PZ73|F9PZ73_9STRE SseB domain-containing protein OS=Streptococcus infantis SK970 OX=1035189 GN=HMPREF9954_0345 PE=4 SV=1
MM1 pKa = 7.35KK2 pKa = 9.43RR3 pKa = 11.84TYY5 pKa = 10.24QPSKK9 pKa = 9.82LRR11 pKa = 11.84RR12 pKa = 11.84ARR14 pKa = 11.84KK15 pKa = 8.57HH16 pKa = 4.75GFRR19 pKa = 11.84NRR21 pKa = 11.84MSTKK25 pKa = 9.22NGRR28 pKa = 11.84RR29 pKa = 11.84VLAARR34 pKa = 11.84RR35 pKa = 11.84RR36 pKa = 11.84KK37 pKa = 8.87GRR39 pKa = 11.84KK40 pKa = 8.75VLAAA44 pKa = 4.31
MM1 pKa = 7.35KK2 pKa = 9.43RR3 pKa = 11.84TYY5 pKa = 10.24QPSKK9 pKa = 9.82LRR11 pKa = 11.84RR12 pKa = 11.84ARR14 pKa = 11.84KK15 pKa = 8.57HH16 pKa = 4.75GFRR19 pKa = 11.84NRR21 pKa = 11.84MSTKK25 pKa = 9.22NGRR28 pKa = 11.84RR29 pKa = 11.84VLAARR34 pKa = 11.84RR35 pKa = 11.84RR36 pKa = 11.84KK37 pKa = 8.87GRR39 pKa = 11.84KK40 pKa = 8.75VLAAA44 pKa = 4.31
Molecular weight: 5.27 kDa
Isoelectric point according different methods:
Peptides (in silico digests for buttom-up proteomics)
Below you can find in silico digests of the whole proteome with Trypsin, Chymotrypsin, Trypsin+LysC, LysN, ArgC proteases suitable for different mass spec machines.| Try ESI |
 |
|---|
| ChTry ESI |
 |
|---|
| ArgC ESI |
 |
|---|
| LysN ESI |
 |
|---|
| TryLysC ESI |
 |
|---|
| Try MALDI |
 |
|---|
| ChTry MALDI |
 |
|---|
| ArgC MALDI |
 |
|---|
| LysN MALDI |
 |
|---|
| TryLysC MALDI |
 |
|---|
| Try LTQ |
 |
|---|
| ChTry LTQ |
 |
|---|
| ArgC LTQ |
 |
|---|
| LysN LTQ |
 |
|---|
| TryLysC LTQ |
 |
|---|
| Try MSlow |
 |
|---|
| ChTry MSlow |
 |
|---|
| ArgC MSlow |
 |
|---|
| LysN MSlow |
 |
|---|
| TryLysC MSlow |
 |
|---|
| Try MShigh |
 |
|---|
| ChTry MShigh |
 |
|---|
| ArgC MShigh |
 |
|---|
| LysN MShigh |
 |
|---|
| TryLysC MShigh |
 |
|---|
General Statistics
Number of major isoforms |
Number of additional isoforms |
Number of all proteins |
Number of amino acids |
Min. Seq. Length |
Max. Seq. Length |
Avg. Seq. Length |
Avg. Mol. Weight |
|---|---|---|---|---|---|---|---|
0 |
564361 |
37 |
2882 |
263.2 |
29.55 |
Amino acid frequency
Ala |
Cys |
Asp |
Glu |
Phe |
Gly |
His |
Ile |
Lys |
Leu |
|---|---|---|---|---|---|---|---|---|---|
7.409 ± 0.061 | 0.543 ± 0.017 |
5.738 ± 0.043 | 7.486 ± 0.063 |
4.486 ± 0.052 | 6.573 ± 0.055 |
1.831 ± 0.024 | 7.06 ± 0.062 |
6.949 ± 0.059 | 9.815 ± 0.09 |
Met |
Asn |
Gln |
Pro |
Arg |
Ser |
Thr |
Val |
Trp |
Tyr |
|---|---|---|---|---|---|---|---|---|---|
2.461 ± 0.029 | 4.499 ± 0.038 |
3.445 ± 0.049 | 4.067 ± 0.043 |
4.035 ± 0.036 | 6.08 ± 0.05 |
5.844 ± 0.081 | 6.987 ± 0.047 |
0.922 ± 0.017 | 3.767 ± 0.042 |
Most of the basic statistics you can see at this page can be downloaded from this CSV file
Proteome-pI is available under Creative Commons Attribution-NoDerivs license, for more details see here
| Reference: Kozlowski LP. Proteome-pI 2.0: Proteome Isoelectric Point Database Update. Nucleic Acids Res. 2021, doi: 10.1093/nar/gkab944 | Contact: Lukasz P. Kozlowski |
